Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).Multiple Linear Regression 1
Lecture 17
Duke University
STA 199 Spring 2026
2026-03-23
While you wait: Participate 📱💻
Which of the following is true about linear regression models?
- The regression line always passes through the origin (x = 0, y = 0).
- The slope indicates the average change in the outcome for a one-unit increase in the predictor.
- The intercept indicates the average value of the predictor when the outcome is 0.
- Least squares regression line minimizes the sum of the vertical distances between the observed and predicted values of the outcome.
Scan the QR code or go HERE. Log in with your Duke NetID.
Do you know what Duke invests in? Neither does Nathan!

Data Manager for ER
- Due: March 28th, 2026 11:59pm;
- Use STA 199 in real-world application;
- Support local Durham community.
Bookbagging this Thursday!

Get great advice about course selection from the old broads in the stats mafia:
- Free boba!
- Thursday 3/26 7pm - 9pm;
- Social Sciences 136
- Instagram: @dukessmu
“I can’t attend lab on March 26”
If you will not be physically present during your team’s presentation in lab this Thursday…
- There are too many of you to make special arrangements;
- There will not be a remote option;
- Your team will present without you;
- You should communicate with your group in advance;
- Everyone on the team gets the same grade on the presentation regardless their attendance;
- We leave it to you to police this. If someone doesn’t communicate, doesn’t contribute, and flakes on the presentation, tear them to shreds in the peer eval.
Recap: simple linear regression
From last time…penguins
A researcher wants to see how body mass varies with flipper length.
outcome: body mass in grams (numerical)
predictor: flipper length in mm (numerical)
Flipper length is easier to measure, so more plausible you would predict body mass based on that and not the other way around.
How do we concisely summarize the association between two variables?
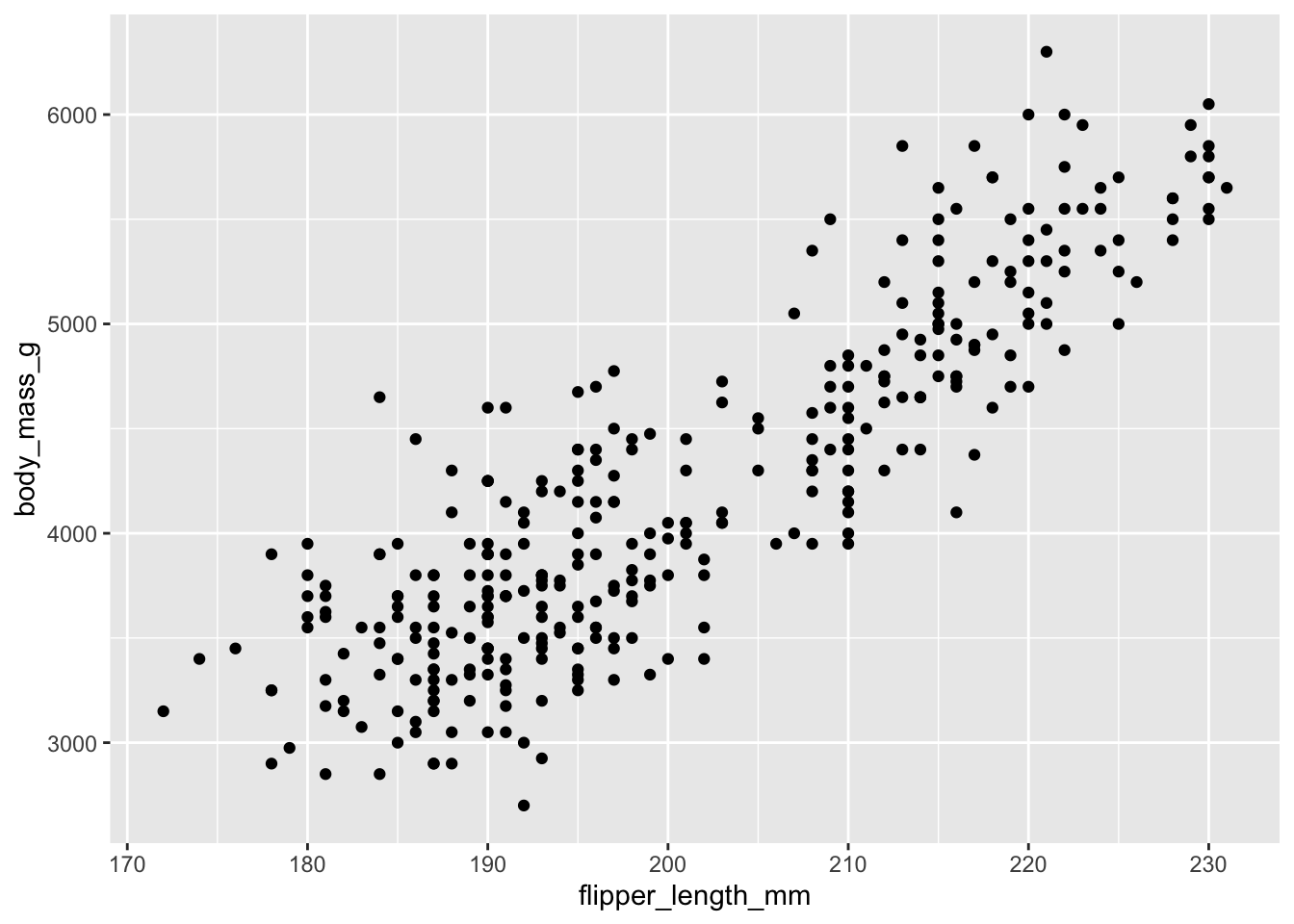
Line of best fit!
ggplot(penguins, aes(x = flipper_length_mm, y = body_mass_g)) +
geom_point() +
geom_smooth(method = "lm")`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_smooth()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).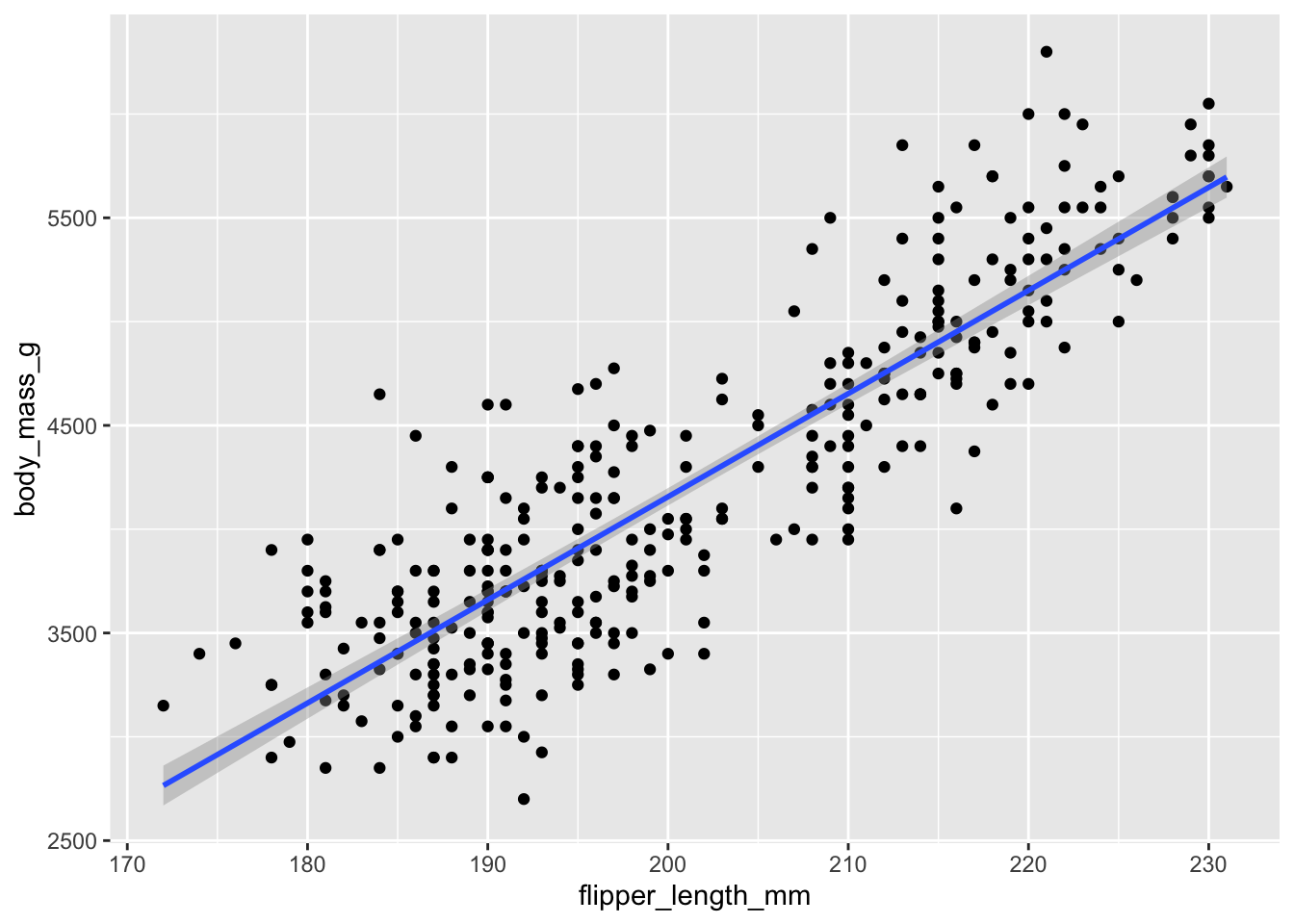
Concise numerical summary: correlation
# A tibble: 1 × 1
r
<dbl>
1 0.871Measures the strength and direction of the linear association between two numerical variables. Strong and positive in this case.
Model fit
# A tibble: 2 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) -5781. 306. -18.9 5.59e- 55
2 flipper_length_mm 49.7 1.52 32.7 4.37e-107\[ \widehat{body~mass}=-5780.83+49.68\times flipper~length \]
\(R^2\) (coefficient of determination)
The fraction of the variation in the response explained by the model. A number between 0 (bad) and 1 (good) that measures goodness-of-fit:
Interpretation
\[ \widehat{body~mass}=-5780.83+49.68\times flipper~length \]
- Intercept: Penguins with 0 flipper length are expected, on average, to weigh -5,781 grams (makes no sense!);
- Slopes: For each additional millimeter of a penguin’s flipper length, the weight of the penguin is expected to be higher, on average, by 49.7 grams.
Prediction
`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_smooth()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).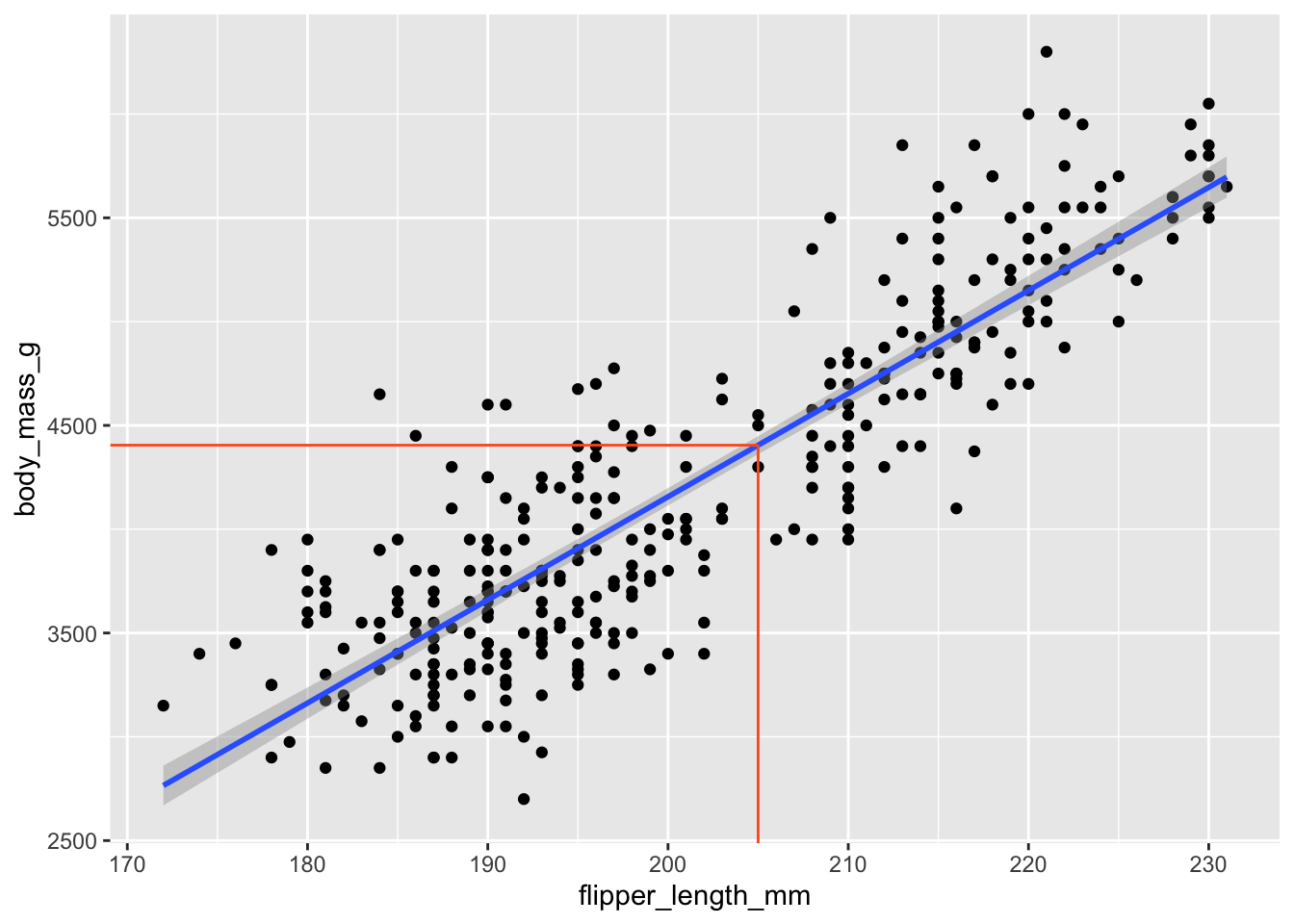
Prediction
# A tibble: 1 × 1
.pred
<dbl>
1 4405.\[ \widehat{body~mass}=-5780.83+49.68\times 205\approx 4404.71 \]
We predict that a penguin whose flipper is 205 mm long will weigh 4404.71 grams on average.
Linear regression with one categorical predictor
New question
A different researcher wants to look at body weight of penguins based on the island they were recorded on. How are the variables involved in this analysis different?
outcome: body mass in grams (numerical)
predictor: island (categorical with three levels)
Participate 📱💻
Determine whether each of the following plot types would be an appropriate choice for visualizing the relationship between body weight and island of penguins.
Scatterplot
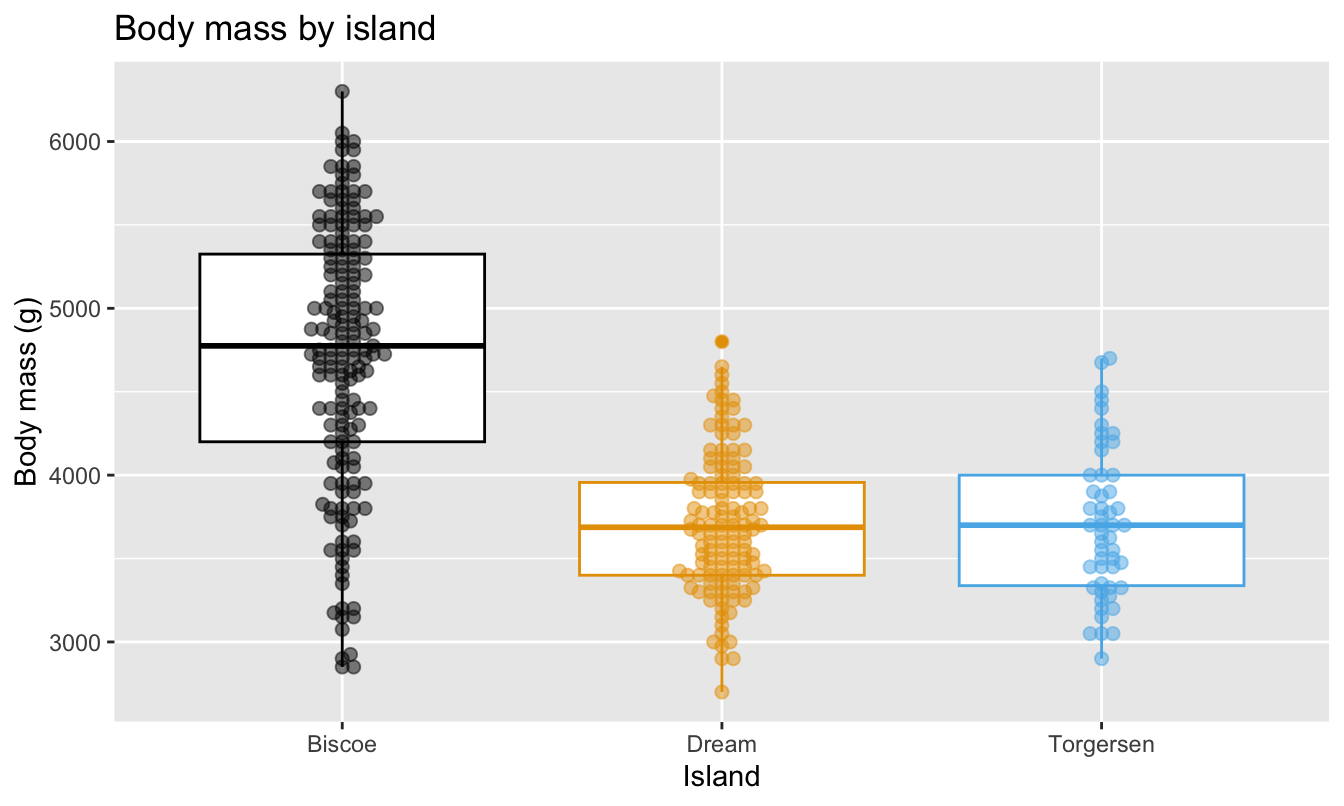
Box plot
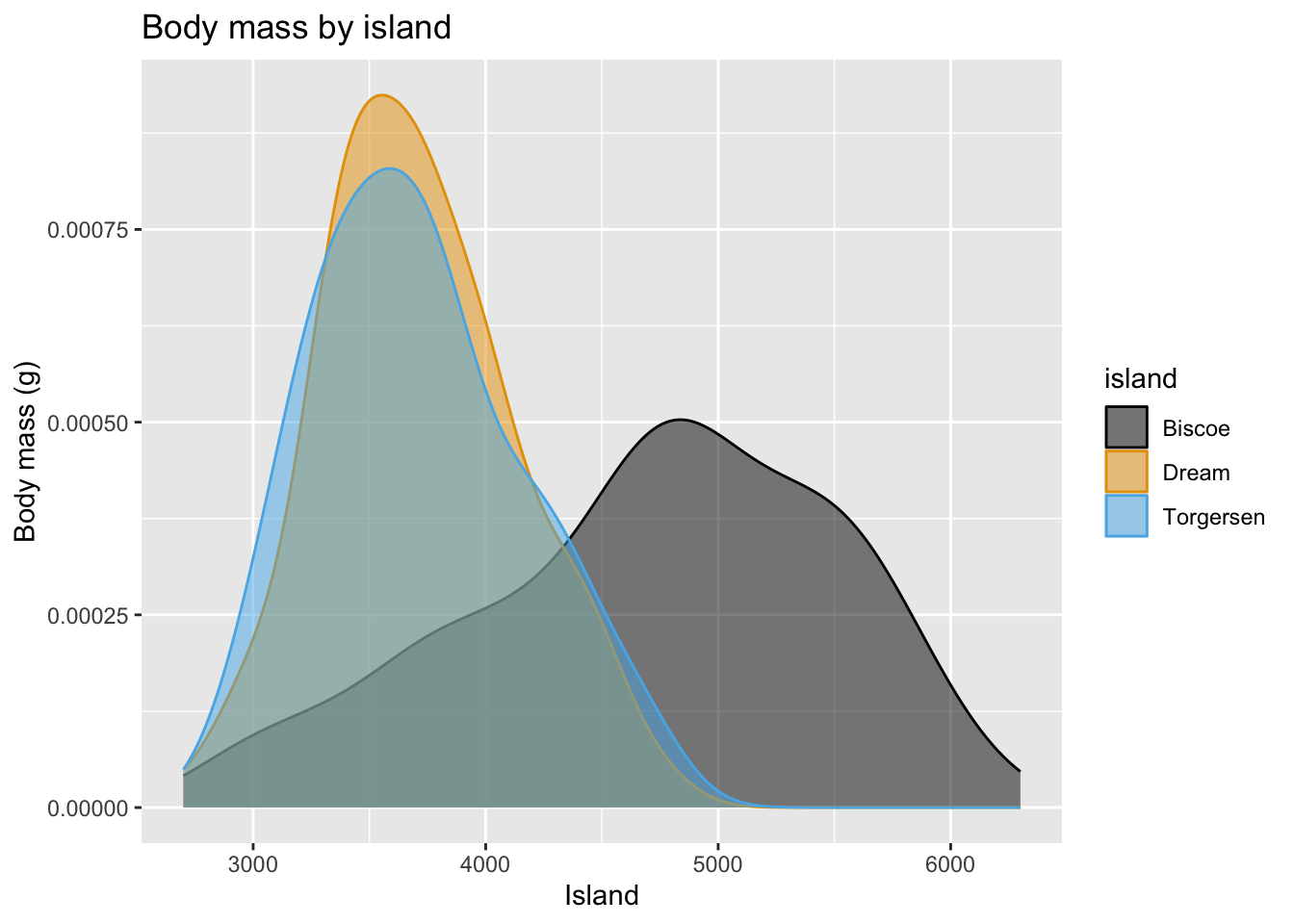
Density plot
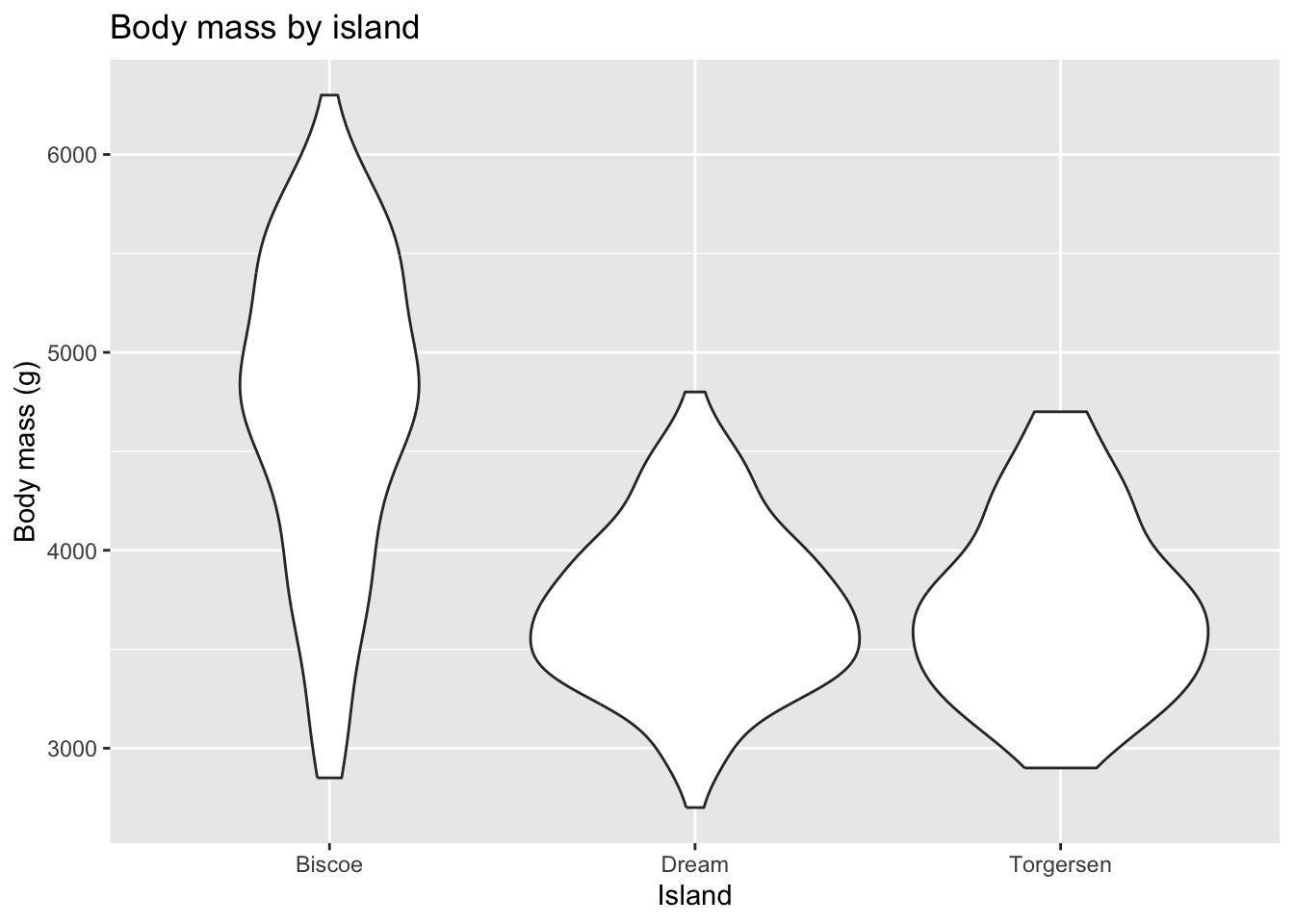
Violin plot
Bar plot
Stacked bar plot
Scan the QR code or go HERE. Log in with your Duke NetID.
Scatterplot: possible, but not so good
Visualize the relationship between body weight and island of penguins. Also calculate the average body weight per island.
Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).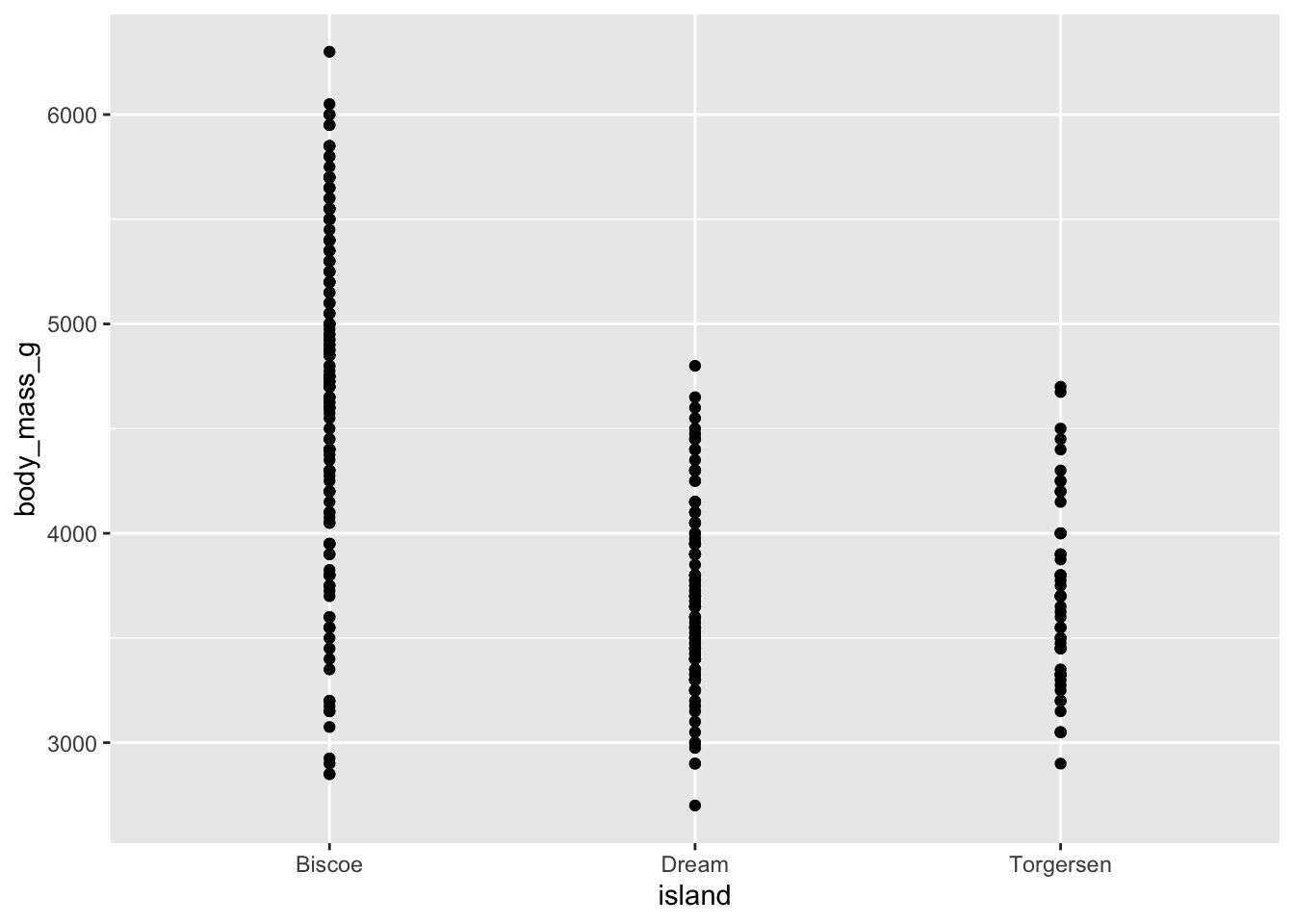
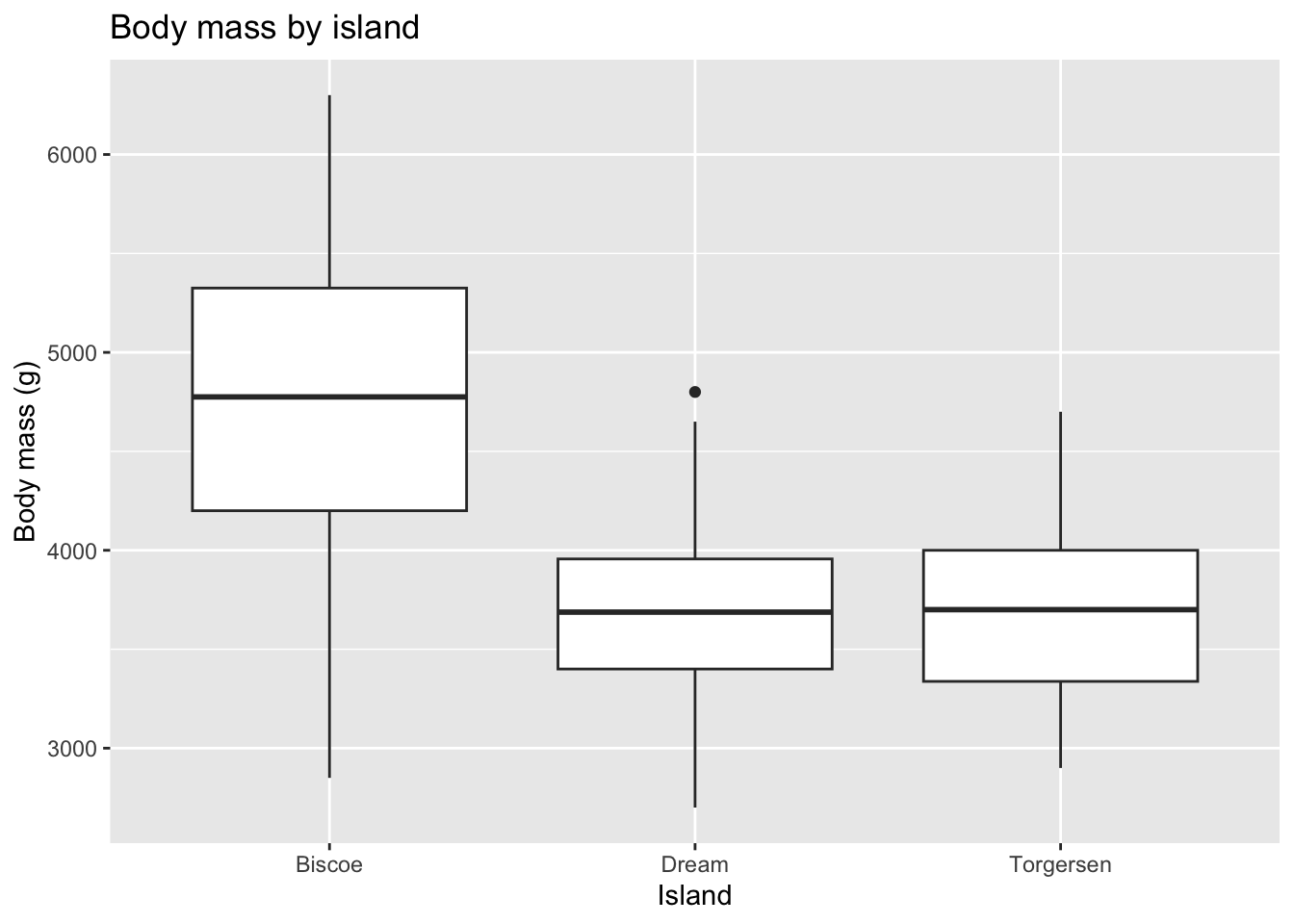
Boxplot
Visualize the relationship between body weight and island of penguins. Also calculate the average body weight per island.
Density plot
Violin plot
Multiple geoms
Model - fit
Fit a linear regression model predicting body weight from island and display the results. Why is Biscoe not on the output?
# A tibble: 3 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 4716. 48.5 97.3 8.93e-250
2 islandDream -1003. 74.2 -13.5 1.42e- 33
3 islandTorgersen -1010. 100. -10.1 4.66e- 21Huh?
Dummy variables
dummy_penguins <- penguins |>
select(body_mass_g, island) |>
arrange(body_mass_g) |>
mutate(
islandDream = if_else(island == "Dream", 1, 0),
islandTorgersen = if_else(island == "Torgersen", 1, 0),
)
dummy_penguins# A tibble: 344 × 4
body_mass_g island islandDream islandTorgersen
<int> <fct> <dbl> <dbl>
1 2700 Dream 1 0
2 2850 Biscoe 0 0
3 2850 Biscoe 0 0
4 2900 Biscoe 0 0
5 2900 Dream 1 0
6 2900 Torgersen 0 1
7 2900 Dream 1 0
8 2925 Biscoe 0 0
9 2975 Dream 1 0
10 3000 Dream 1 0
# ℹ 334 more rowsDummy variables
# A tibble: 3 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 4716. 48.5 97.3 8.93e-250
2 islandDream -1003. 74.2 -13.5 1.42e- 33
3 islandTorgersen -1010. 100. -10.1 4.66e- 21Exactly the same as before!
Model - interpret
\[ \widehat{body~mass} = 4716 - 1003 \times islandDream - 1010 \times islandTorgersen \]
Intercept: Penguins from Biscoe island are expected to weigh, on average, 4,716 grams.
Slope - islandDream: Penguins from Dream island are expected to weigh, on average, 1,003 grams less than those from Biscoe island.
Slope - islandTorgersen: Penguins from Torgersen island are expected to weigh, on average, 1,010 grams less than those from Biscoe island.
Model - predict
What is the predicted body weight of a penguin on Biscoe island? What are the estimated body weights of penguins on Dream and Torgersen islands? Where have we seen these values before?
Model - predict
Calculate the predicted body weights of penguins on Biscoe, Dream, and Torgersen islands by hand.
\[ \widehat{body~mass} = 4716 - 1003 \times islandDream - 1010 \times islandTorgersen \]
- Biscoe: \(\widehat{body~mass} = 4716 - 1003 \times 0 - 1010 \times 0 = 4716\)
- Dream: \(\widehat{body~mass} = 4716 - 1003 \times 1 - 1010 \times 0 = 3713\)
- Torgersen: \(\widehat{body~mass} = 4716 - 1003 \times 0 - 1010 \times 1 = 3706\)
Look familiar?
Models with categorical predictors
When the categorical predictor has many levels, they’re encoded as dummy variables.
The first level of the categorical variable is the baseline level. In a model with one categorical predictor, the intercept is the predicted value of the outcome for the baseline level (x = 0).
Each slope coefficient describes the difference between the predicted value of the outcome for that level of the categorical variable compared to the baseline level.
In general
Predicting continuous outcome \(Y\) using one categorical predictor \(X\) with multiple levels 0, 1, 2, …, \(L\). Create dummy variables for every level except the base level:
\[ cat_l=\begin{cases} 1 & X=l\\ 0 & \text{else} \end{cases} \]
Then fit a regression with multiple dummy predictors:
\[ \hat{Y} = b_0 + b_1 \times cat_1 + b_2 \times cat_2 \ldots + b_L \times cat_L \]
\(b_0\) : the model prediction for a member of the base level;
\(b_1\): how does the prediction change when we move from the base level to level 1?
\(b_2\): how does the prediction change when we move from the base level to level 2?
etc
We’re not animals. We have technology!
The computer handles all of this for you, but you need to understand the details so you code and interpret it correctly.
Changing the baseline level
By default, R uses the first level of a categorical variable as the baseline level. this is often the first alphabetically, but make sure you check!
We can change the baseline level by reordering the levels of the categorical variable.
Participate 📱💻
Scan the QR code or go HERE. Log in with your Duke NetID.
Participate 📱💻
What is the baseline level of island in the following model??
# A tibble: 3 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 3706. 87.7 42.3 3.64e-137
2 islandBiscoe 1010. 100. 10.1 4.66e- 21
3 islandDream 6.53 104. 0.0627 9.50e- 1- Biscoe
- Dream
- Torgersen
- Cannot tell from the output
Scan the QR code or go HERE. Log in with your Duke NetID.
Linear regression with one numerical and one categorical predictor
Three variables on one 2D plot
Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).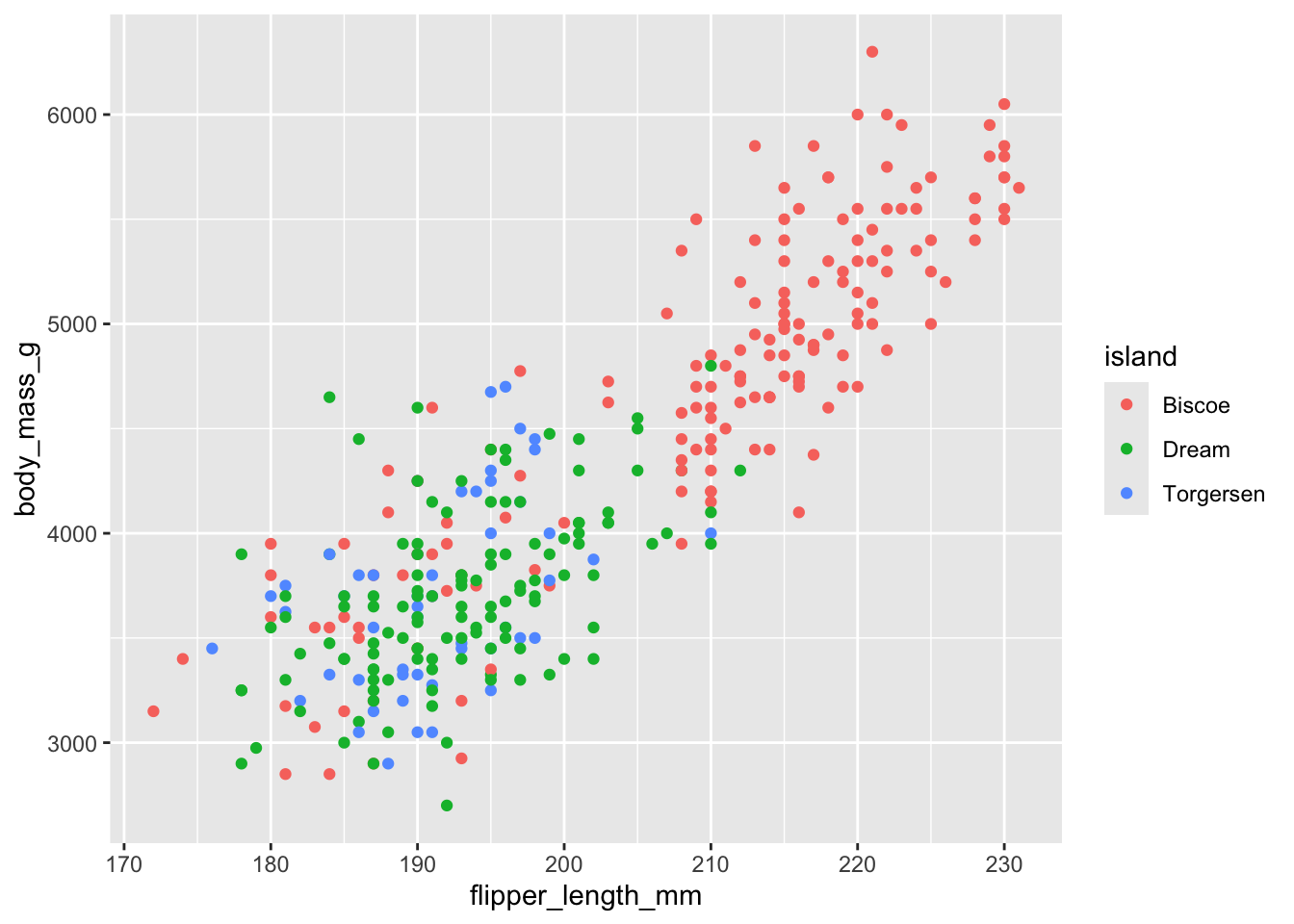
How do we predict using more than one predictor?
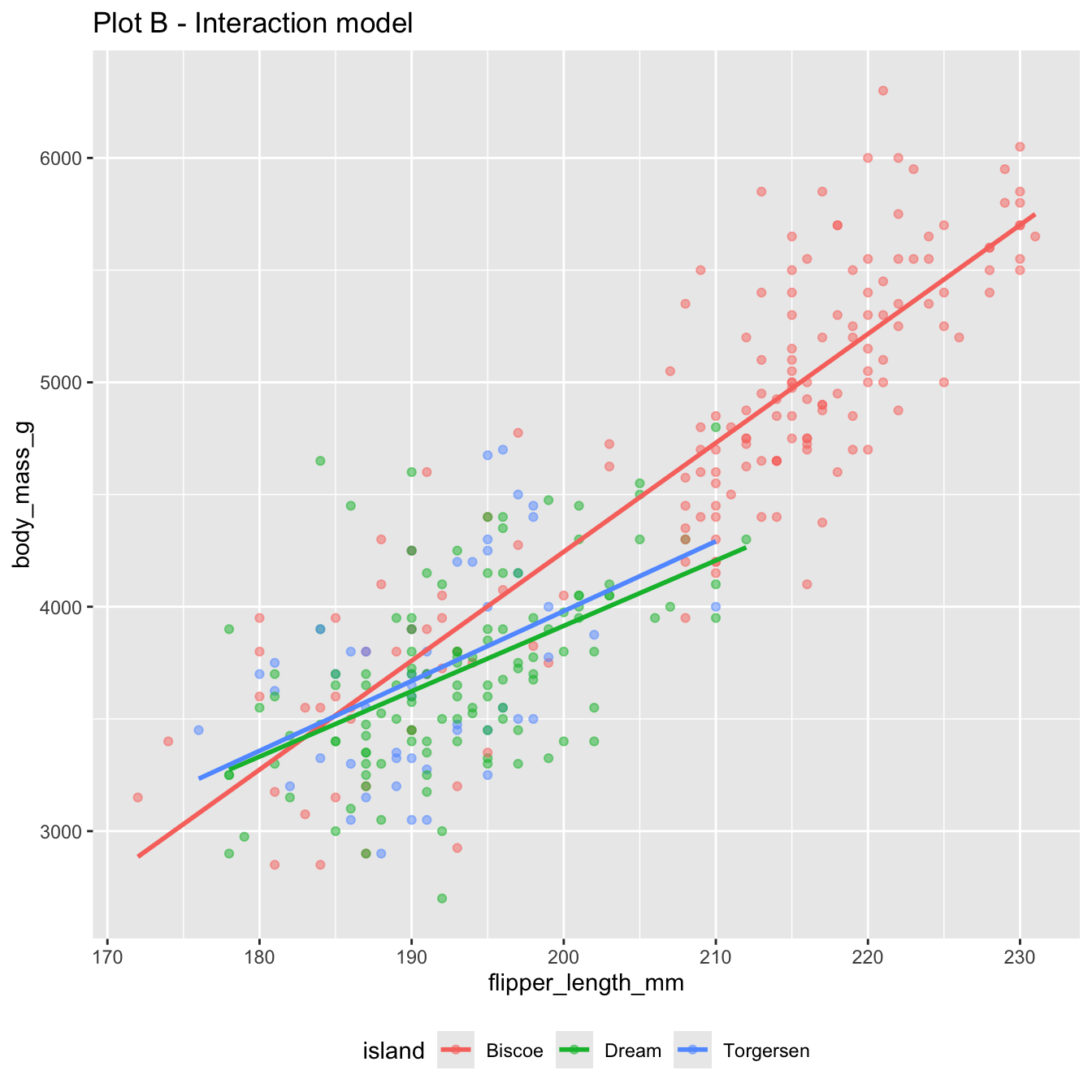
Both of these models use flipper_length_mm and island to predict body_mass_g:
`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_smooth()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range (`stat_smooth()`).
Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).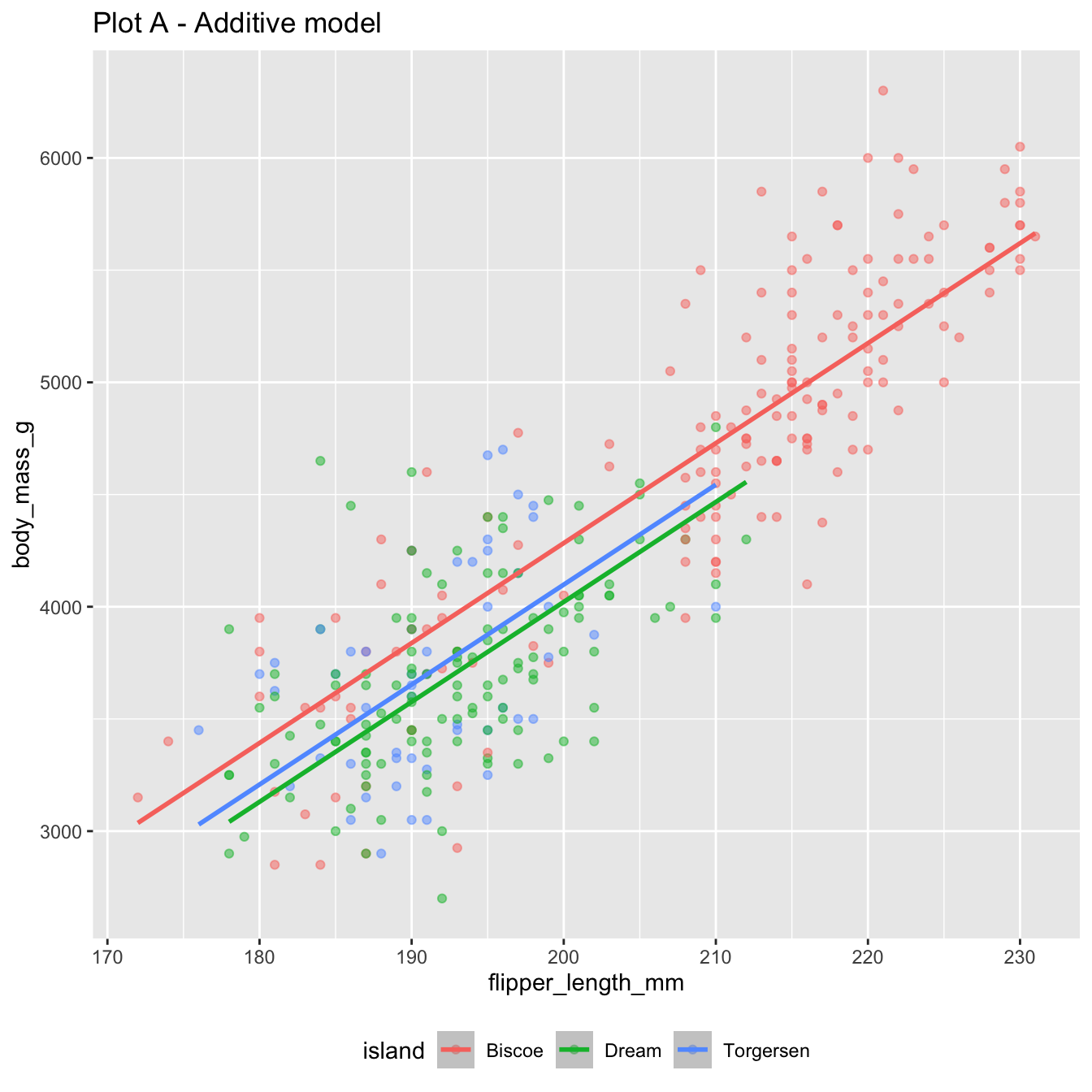
The additive model: parallel lines, one for each island
`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_smooth()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).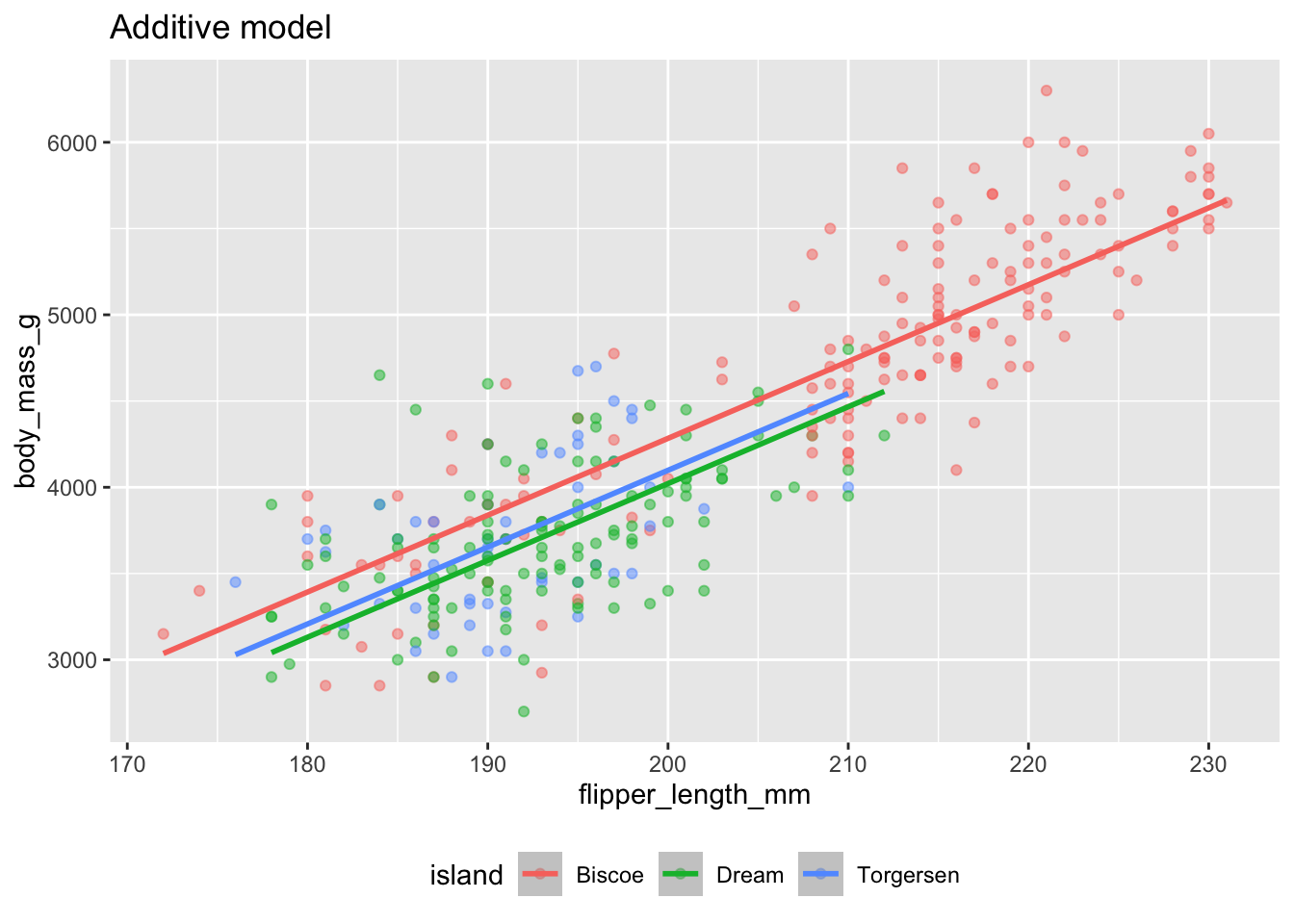
The additive model: parallel lines, one for each island
bm_fl_island_fit <- linear_reg() |>
fit(body_mass_g ~ flipper_length_mm + island, data = penguins)
tidy(bm_fl_island_fit)# A tibble: 4 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) -4625. 392. -11.8 4.29e-27
2 flipper_length_mm 44.5 1.87 23.9 1.65e-74
3 islandDream -262. 55.0 -4.77 2.75e- 6
4 islandTorgersen -185. 70.3 -2.63 8.84e- 3\[ \begin{aligned} \widehat{body~mass} = -4625 &+ 44.5 \times flipper~length \\ &- 262 \times Dream \\ &- 185 \times Torgersen \end{aligned} \]
Where do the three lines come from?
\[ \begin{aligned} \widehat{body~mass} = -4625 &+ 44.5 \times flipper~length \\ &- 262 \times Dream \\ &- 185 \times Torgersen \end{aligned} \]
If penguin is from Biscoe, Dream = 0 and Torgersen = 0:
\[ \begin{aligned} \widehat{body~mass} = -4625 &+ 44.5 \times flipper~length \end{aligned} \]
If penguin is from Dream, Dream = 1 and Torgersen = 0:
\[ \begin{aligned} \widehat{body~mass} = -4887 &+ 44.5 \times flipper~length \end{aligned} \]
If penguin is from Torgersen, Dream = 0 and Torgersen = 1:
\[ \begin{aligned} \widehat{body~mass} = -4810 &+ 44.5 \times flipper~length \end{aligned} \]
Either way, same slope, so the lines are parallel.
The interaction model: different lines for each island
Code
`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_smooth()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).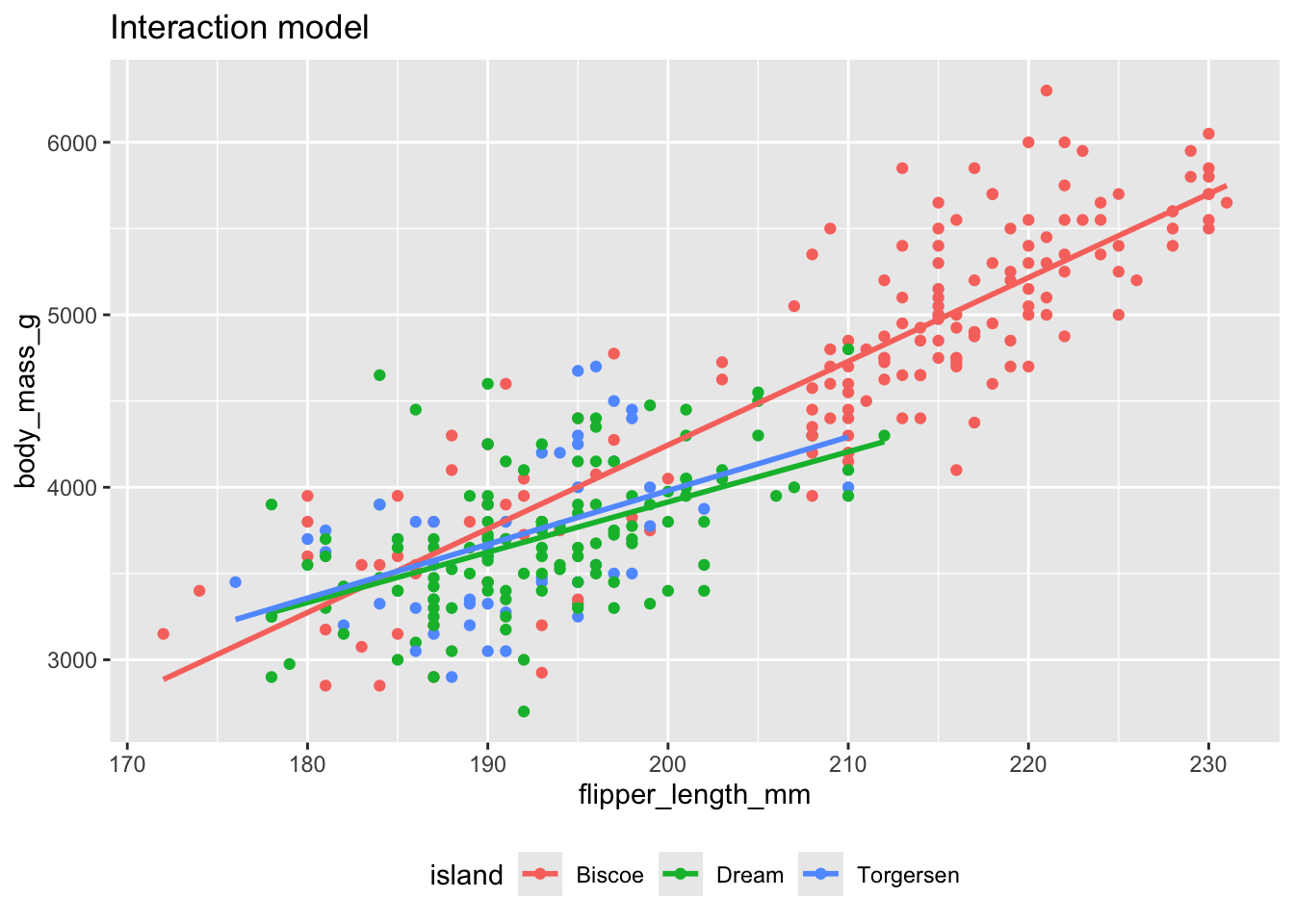
The interaction model: different lines for each island
bm_fl_island_int_fit <- linear_reg() |>
fit(body_mass_g ~ flipper_length_mm * island, data = penguins)
tidy(bm_fl_island_int_fit) |> select(term, estimate)# A tibble: 6 × 2
term estimate
<chr> <dbl>
1 (Intercept) -5464.
2 flipper_length_mm 48.5
3 islandDream 3551.
4 islandTorgersen 3218.
5 flipper_length_mm:islandDream -19.4
6 flipper_length_mm:islandTorgersen -17.4\[ \begin{aligned} \widehat{body~mass} = -5464 &+ 48.5 \times flipper~length \\ &+ 3551 \times Dream \\ &+ 3218 \times Torgersen \\ &- 19.4 \times flipper~length*Dream \\ &- 17.4 \times flipper~length*Torgersen \end{aligned} \]
Where do the three lines come from?
\[ \begin{aligned} \small\widehat{body~mass} = -5464 &+ 48.5 \times flipper~length \\ &+ 3551 \times Dream \\ &+ 3218 \times Torgersen \\ &- 19.4 \times flipper~length*Dream \\ &- 17.4 \times flipper~length*Torgersen \end{aligned} \]
If penguin is from Biscoe, Dream = 0 and Torgersen = 0:
\[ \begin{aligned} \widehat{body~mass} = -5464 &+ 48.5 \times flipper~length \end{aligned} \]
If penguin is from Dream, Dream = 1 and Torgersen = 0:
\[ \begin{aligned} \widehat{body~mass} &= (-5464 + 3551) + (48.5-19.4) \times flipper~length\\ &=-1913+29.1\times flipper~length. \end{aligned} \]
Prediction on Biscoe island
`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_smooth()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).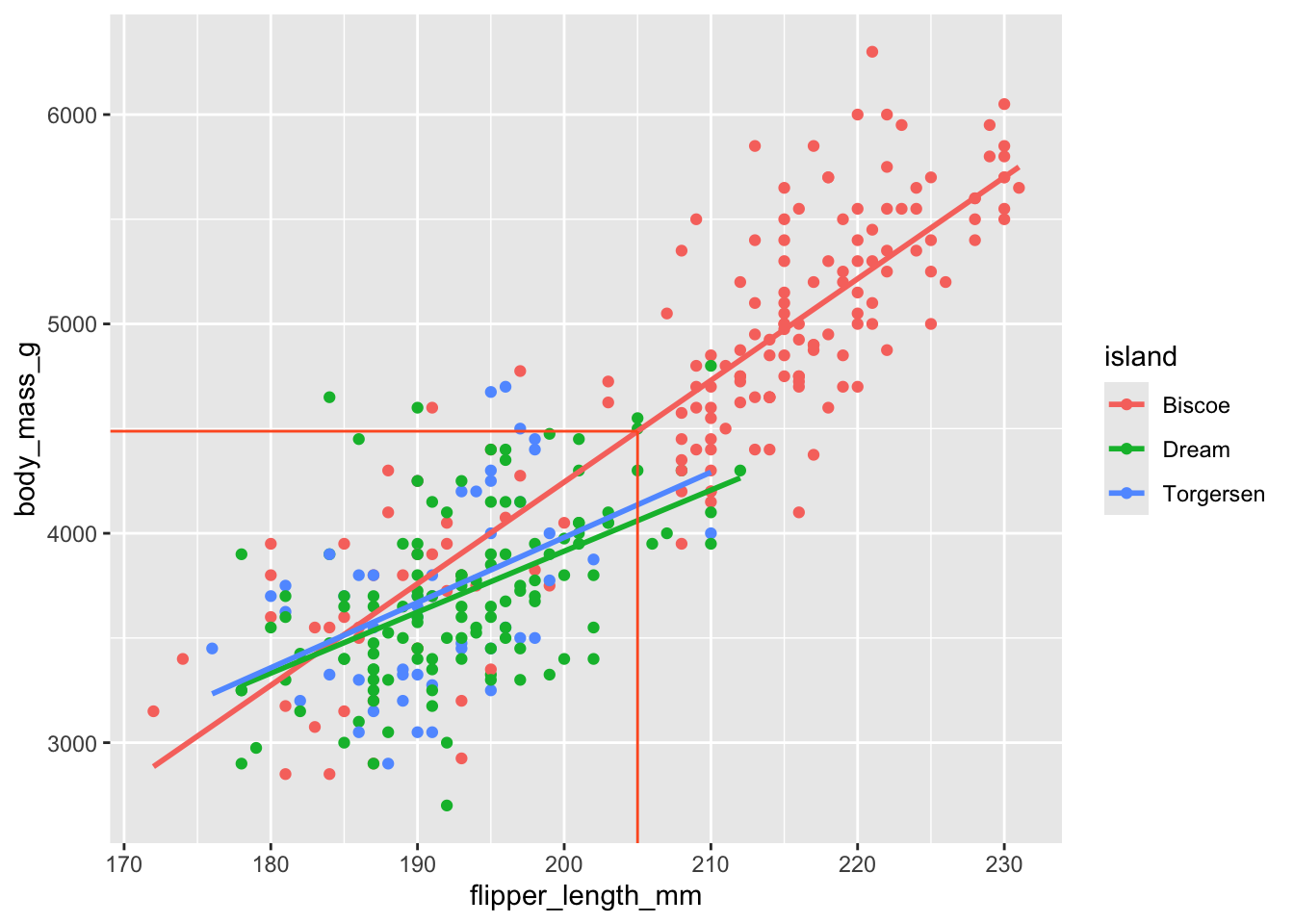
Prediction on Biscoe island
new_penguin <- tibble(
flipper_length_mm = 205,
island = "Biscoe"
)
predict(bm_fl_island_int_fit, new_data = new_penguin)# A tibble: 1 × 1
.pred
<dbl>
1 4488.\[ \widehat{body~mass} = -5464 + 48.5 \times 205. \]
Prediction on Dream island
`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_smooth()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).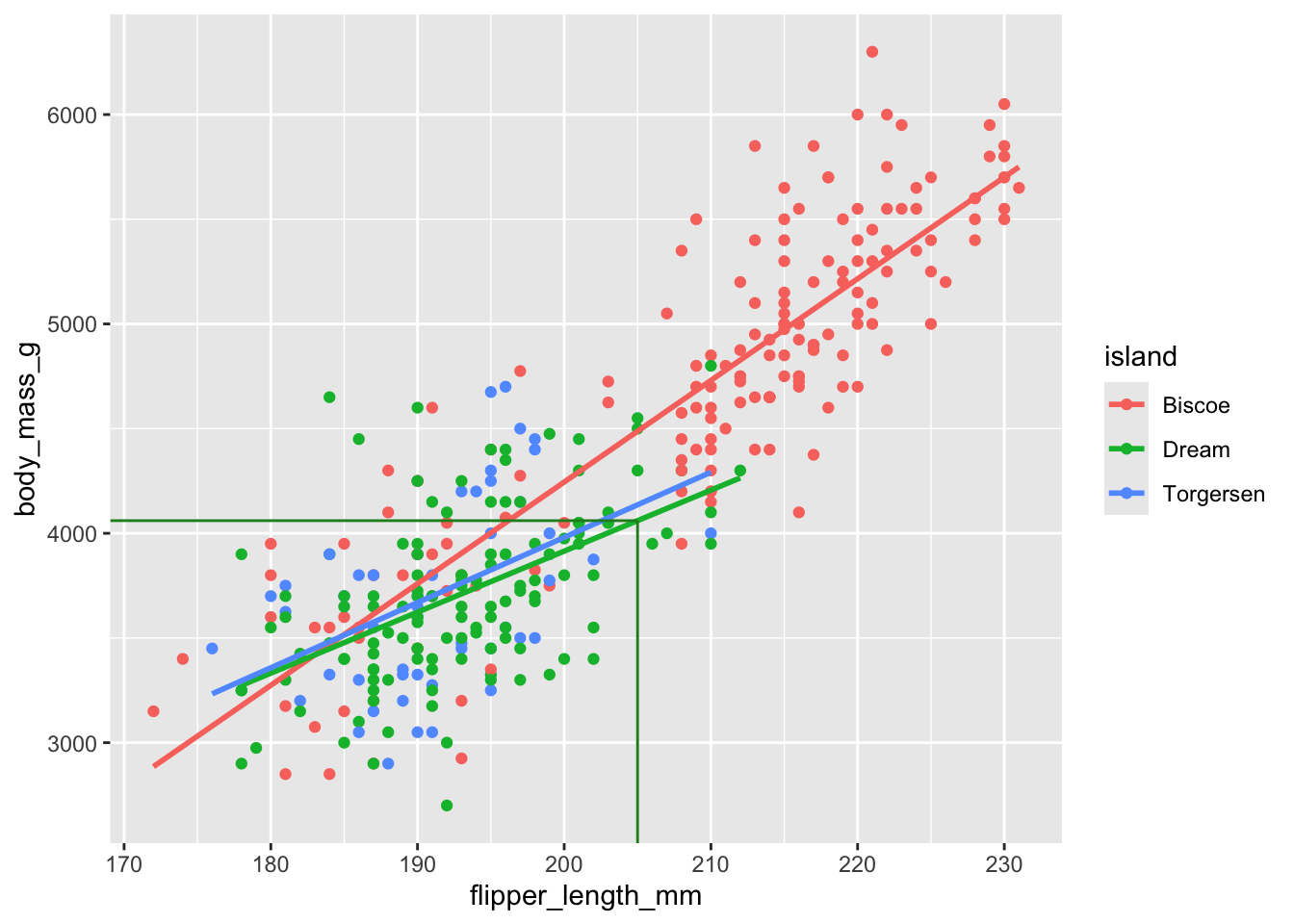
Prediction on Dream island
new_penguin <- tibble(
flipper_length_mm = 205,
island = "Dream"
)
predict(bm_fl_island_int_fit, new_data = new_penguin)# A tibble: 1 × 1
.pred
<dbl>
1 4060.\[ \widehat{body~mass} = (-5464 + 3551) + (48.5 - 19.4) \times 205. \]
Prediction on Torgersen island
`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_smooth()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).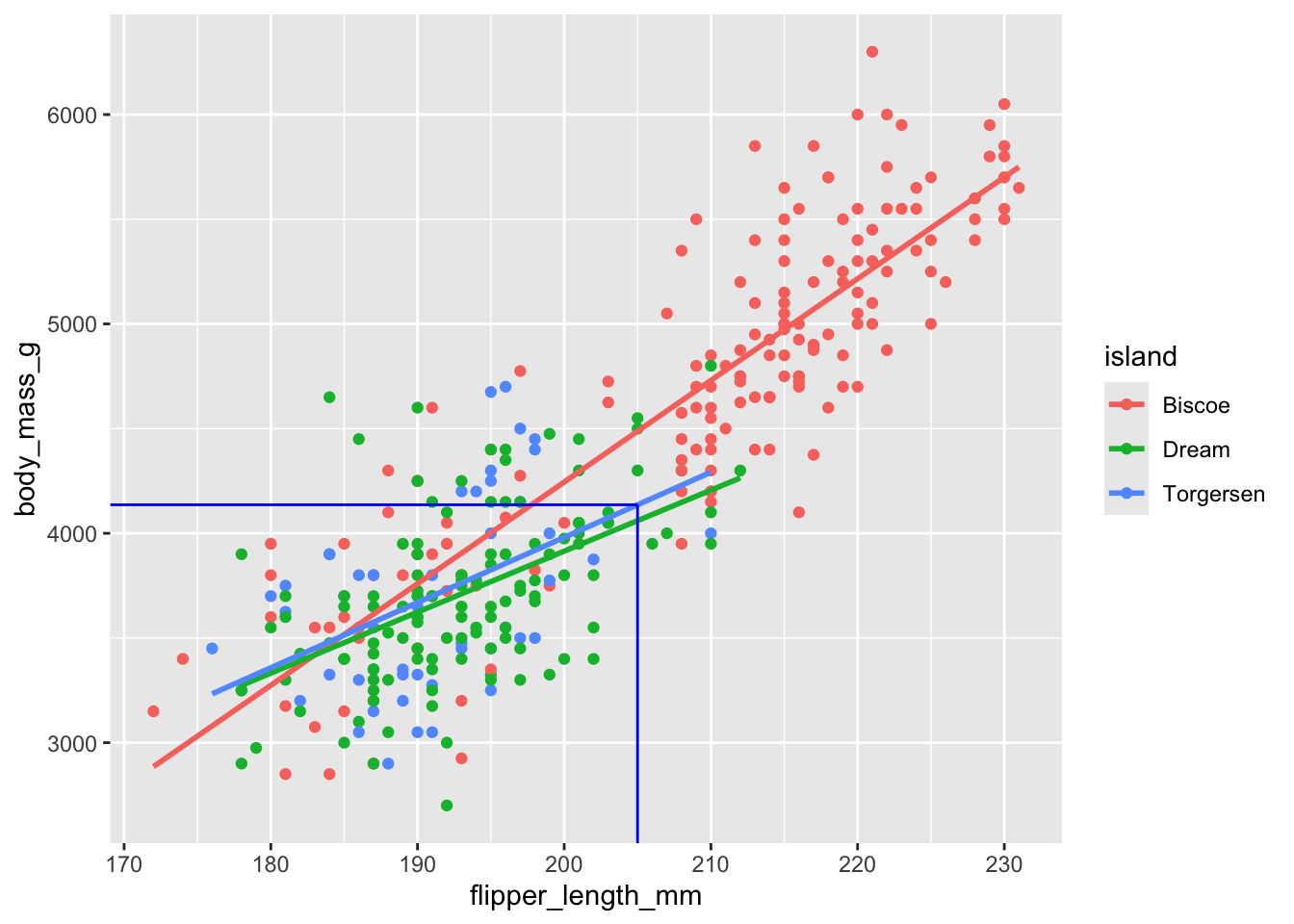
Prediction on Torgersen island
new_penguin <- tibble(
flipper_length_mm = 205,
island = "Torgersen"
)
predict(bm_fl_island_int_fit, new_data = new_penguin)# A tibble: 1 × 1
.pred
<dbl>
1 4136.\[ \widehat{body~mass} = (-5464 + 3218) + (48.5 - 17.4) \times 205. \]
Linear regression with many numerical predictors
Multiple numerical predictors
bm_fl_bl_fit <- linear_reg() |>
fit(body_mass_g ~ flipper_length_mm + bill_length_mm, data = penguins)
tidy(bm_fl_bl_fit)# A tibble: 3 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) -5737. 308. -18.6 7.80e-54
2 flipper_length_mm 48.1 2.01 23.9 7.56e-75
3 bill_length_mm 6.05 5.18 1.17 2.44e- 1\[ \small\widehat{body~mass}=-5736+48.1\times flipper~length+6\times bill~length \]
Interpretation
\[ \small\widehat{body~mass}=-5736+48.1\times flipper~length+6\times bill~length \]
Interpretations:
- We predict that the body mass of a penguin with zero flipper length and zero bill length will be -5736 grams, on average (makes no sense);
- Holding all other variables constant, for every additional millimeter in flipper length, we expect the body mass of penguins to be higher, on average, by 48.1 grams.
- Holding all other variables constant, for every additional millimeter in bill length, we expect the body mass of penguins to be higher, on average, by 6 grams.
Prediction
new_penguin <- tibble(
flipper_length_mm = 200,
bill_length_mm = 45
)
predict(bm_fl_bl_fit, new_data = new_penguin)# A tibble: 1 × 1
.pred
<dbl>
1 4164.\[ \widehat{body~mass}=-5736+48.1\times 200+6\times 45 \]
Picture? It’s not pretty…
2 predictors + 1 response = 3 dimensions. Ick!
Warning: Unknown or uninitialised column: `color`.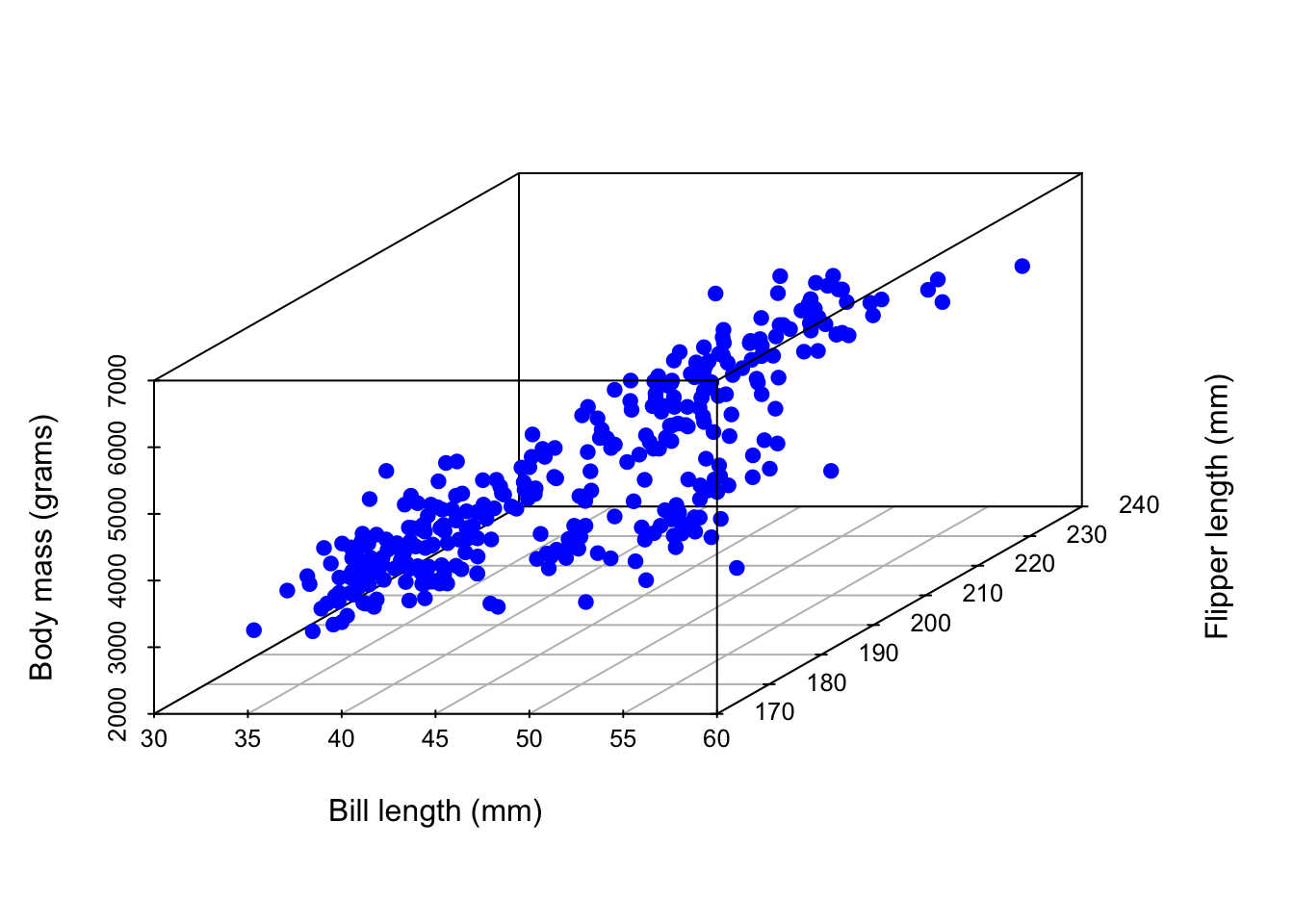
Picture? It’s not pretty…
Instead of a line of best fit, it’s a plane of best fit. Double ick!
Warning: Unknown or uninitialised column: `color`.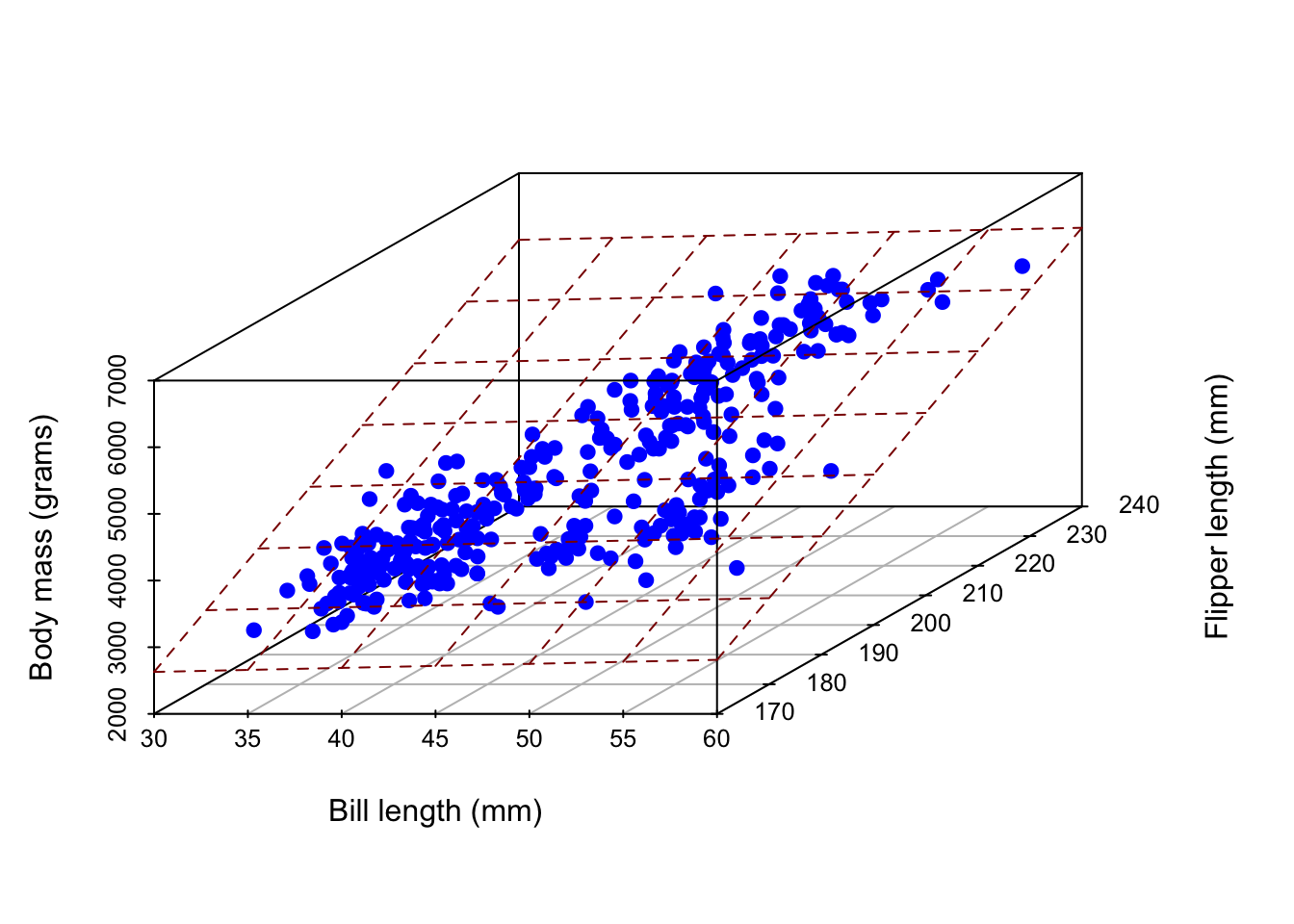
Multiple linear regression in general
Population model
Multiple linear regression captures the relationship between a numerical outcome \(Y\) and many numerical predictors \(X_1\), \(X_2\), …, \(X_p\):
\[\Large{Y = \beta_0 + \beta_1 X_1 + \beta_2 X_2 + ...+ \beta_p X_p+\epsilon}\]
- \(\beta_0\): True intercept of the relationship between the \(X_j\) and \(Y\)
- \(\beta_j\): True slope of the relationship between \(X_j\) and \(Y\)
- \(\epsilon\): Error (residual)
The model with the greek letters and the error term is the “true,” idealized, population relationship that we could access if we had infinite amounts of perfect data. But we don’t, so we have to settle for…
Estimated model
\[\Large{\hat{Y} = b_0 + b_1 X_1 + b_2 X_2 + ... + b_pX_p}\]
- \(b_j\): Estimated slope of the relationship between \(X_j\) and \(Y\);
- \(b_0\): your prediction for \(Y\) if all the predictors equal zero;
- No error term!
This is your best guess at the true regression function based on the noisy, meager, imperfect data you actually have access to. We still compute the \(b_j\) using the principle of least squares: pick the estimates that make the sum of squared residuals as small as possible.
Next time
Today we saw multiple models that are all attempting to do the same thing: predict body mass.
- Which one is “best”?
- What does “best” even mean?