`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_smooth()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).Lecture 18
Duke University
STA 199 Spring 2026
2026-03-25

Get great advice about course selection from the old broads in the stats mafia:
presentations.pdf is up-to-date and your website is working before 8AM tomorrow.Multiple linear regression captures the relationship between a numerical outcome \(Y\) and many numerical predictors \(X_1\), \(X_2\), …, \(X_p\):
\[\Large{Y = \beta_0 + \beta_1 X_1 + \beta_2 X_2 + ...+ \beta_p X_p+\epsilon}\]
The model with the greek letters and the error term is the “true,” idealized, population relationship that we could access if we had infinite amounts of perfect data. But we don’t, so we have to settle for…
\[\Large{\hat{Y} = b_0 + b_1 X_1 + b_2 X_2 + ... + b_pX_p}\]
This is your best guess at the true regression function based on the noisy, meager, imperfect data you actually have access to. We still compute the \(b_j\) using the principle of least squares: pick the estimates that make the sum of squared residuals as small as possible.
`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_smooth()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).# A tibble: 2 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) -5781. 306. -18.9 5.59e- 55
2 flipper_length_mm 49.7 1.52 32.7 4.37e-107\[ \widehat{body~mass}=-5780.83+49.68\times flipper~length \]
`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_smooth()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).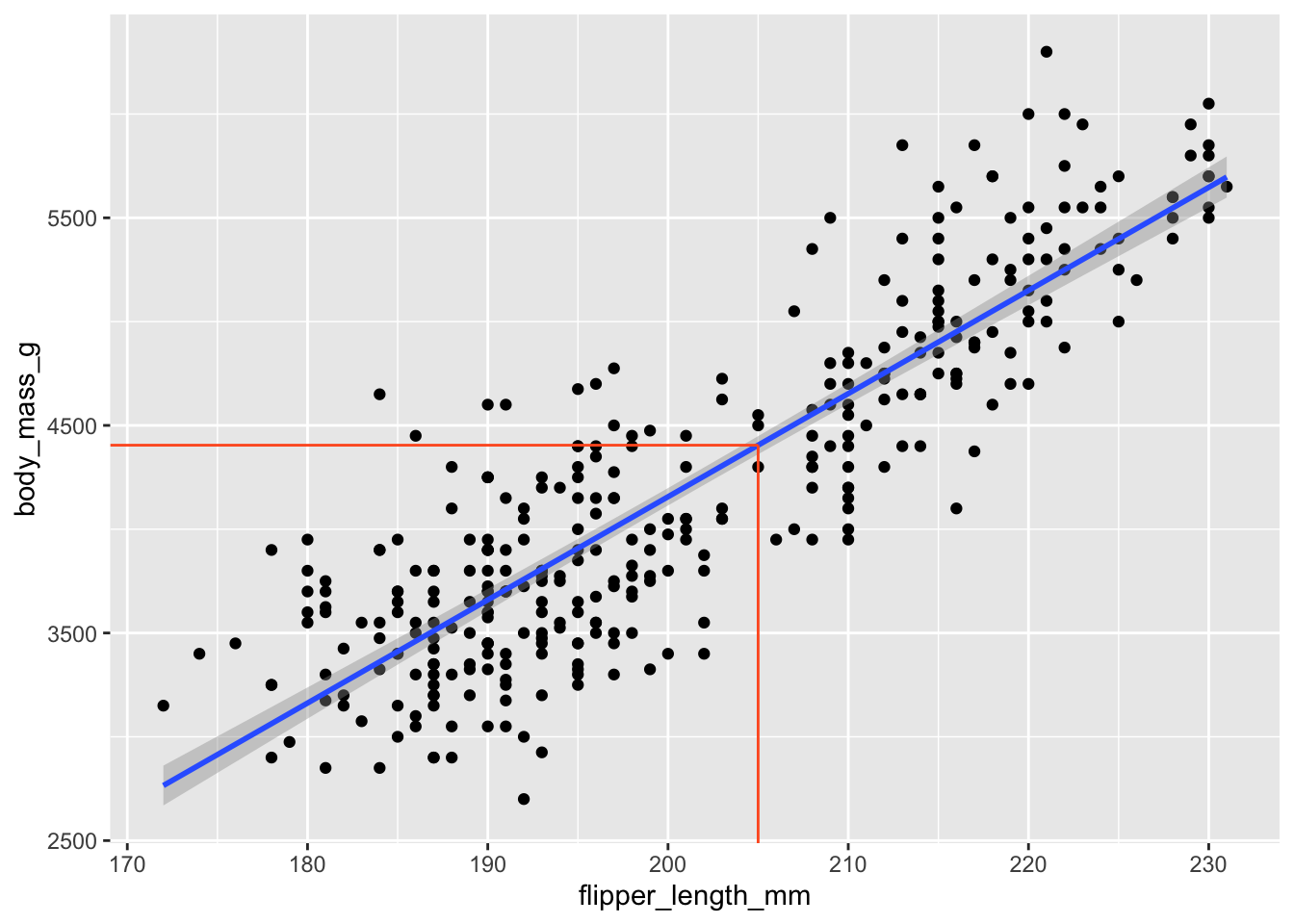
Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_boxplot()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).This actually gets treated as multiple binary (0/1) predictors:
# A tibble: 3 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 4716. 48.5 97.3 8.93e-250
2 islandDream -1003. 74.2 -13.5 1.42e- 33
3 islandTorgersen -1010. 100. -10.1 4.66e- 21\[ \begin{aligned} \widehat{body~mass} = 4716 &- 1003 \times islandDream \\ &- 1010 \times islandTorgersen \end{aligned} \]
It’s just a concise way of summarizing the within-group means of the response variable:
`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_smooth()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range (`stat_smooth()`).
Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).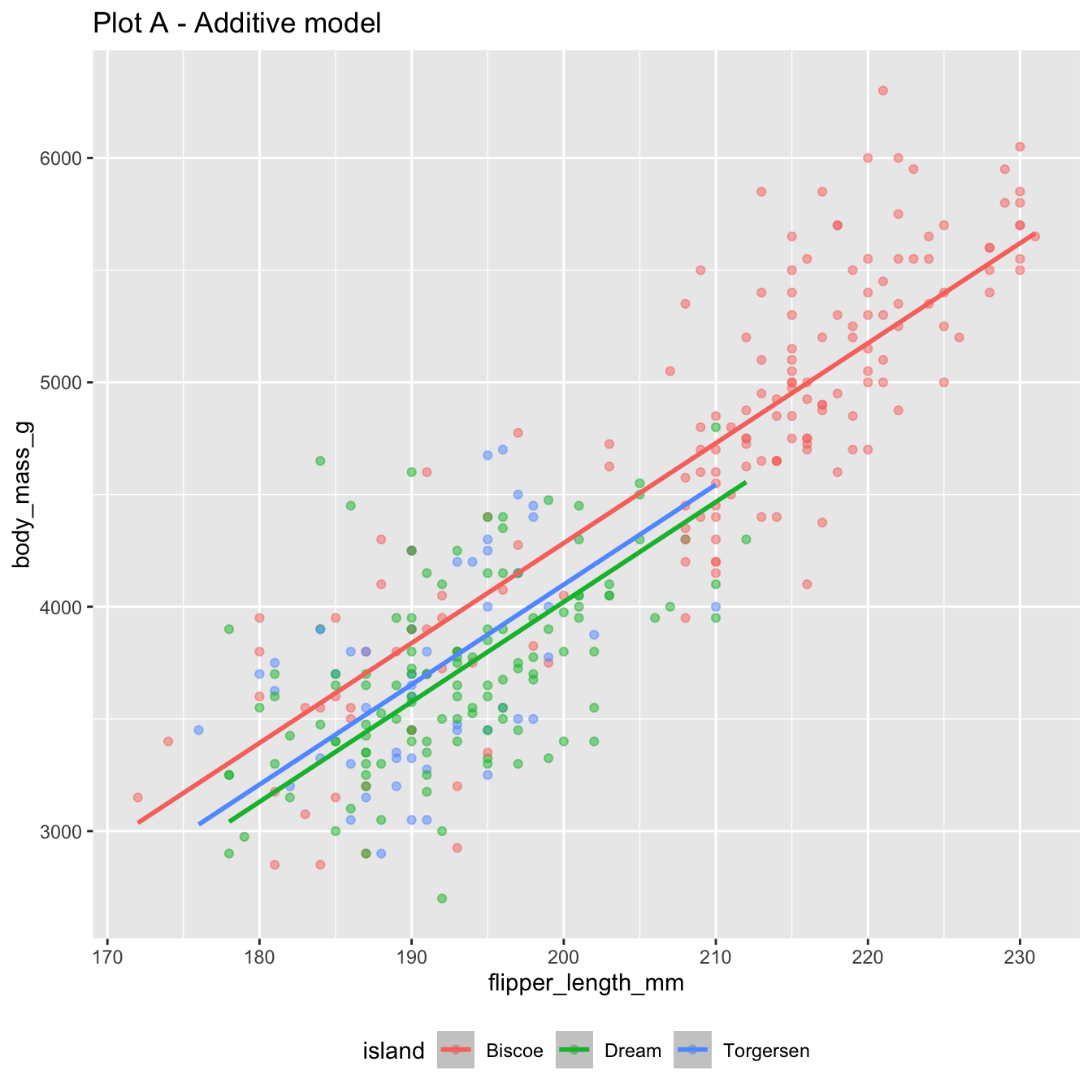
The additive model:
bm_fl_island_fit <- linear_reg() |>
fit(body_mass_g ~ flipper_length_mm + island, data = penguins)
tidy(bm_fl_island_fit)# A tibble: 4 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) -4625. 392. -11.8 4.29e-27
2 flipper_length_mm 44.5 1.87 23.9 1.65e-74
3 islandDream -262. 55.0 -4.77 2.75e- 6
4 islandTorgersen -185. 70.3 -2.63 8.84e- 3\[ \begin{aligned} \widehat{body~mass} = -4625 &+ 44.5 \times flipper~length \\ &- 262 \times Dream \\ &- 185 \times Torgersen \end{aligned} \]
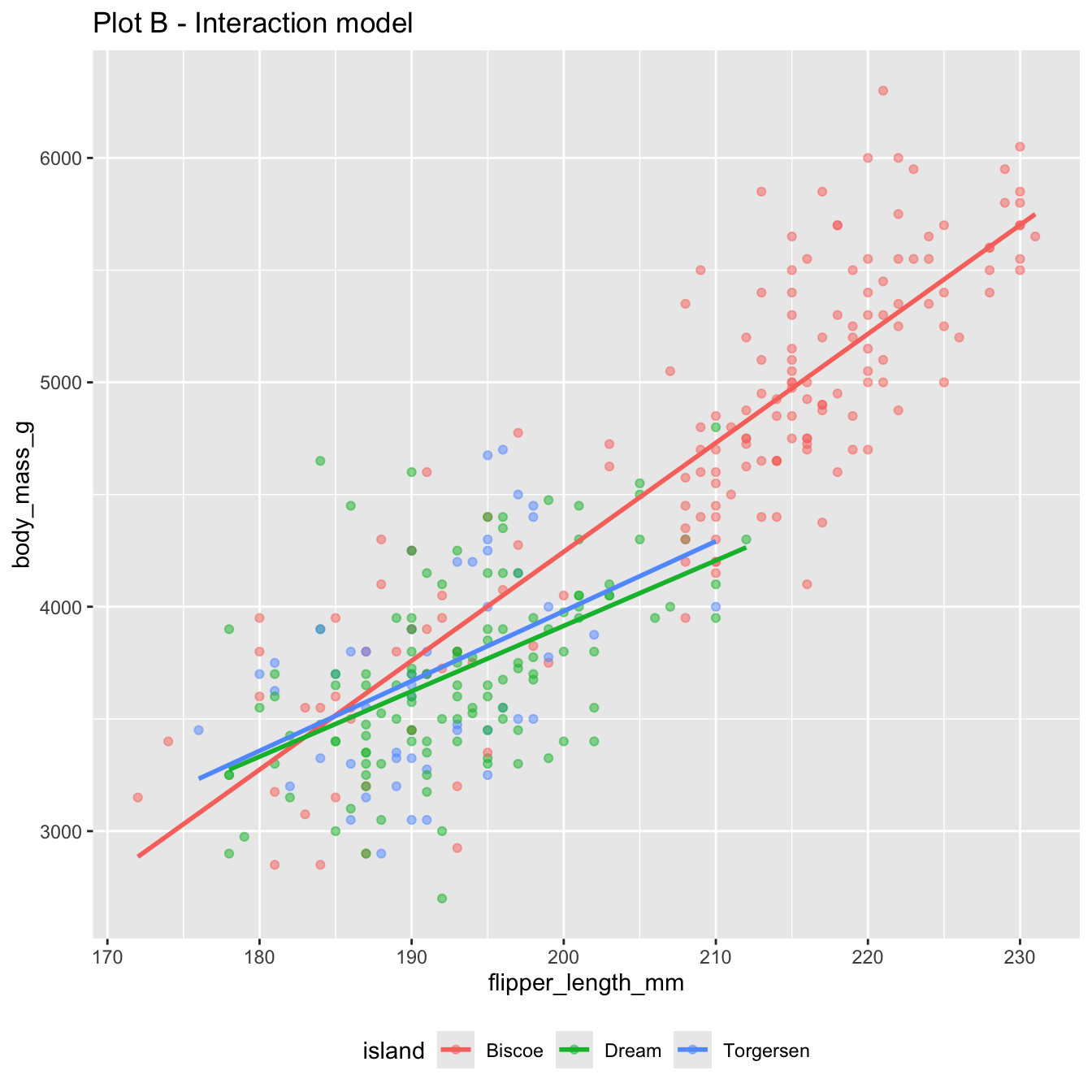
Change one character in the code, and you get the interaction model:
bm_fl_island_int_fit <- linear_reg() |>
fit(body_mass_g ~ flipper_length_mm * island, data = penguins)
tidy(bm_fl_island_int_fit) |> select(term, estimate)# A tibble: 6 × 2
term estimate
<chr> <dbl>
1 (Intercept) -5464.
2 flipper_length_mm 48.5
3 islandDream 3551.
4 islandTorgersen 3218.
5 flipper_length_mm:islandDream -19.4
6 flipper_length_mm:islandTorgersen -17.4\[ \begin{aligned} \widehat{body~mass} = -5464 &+ 48.5 \times flipper~length \\ &+ 3551 \times Dream \\ &+ 3218 \times Torgersen \\ &- 19.4 \times flipper~length*Dream \\ &- 17.4 \times flipper~length*Torgersen \end{aligned} \]
Same code with a different model fit:
2 predictors + 1 response = 3 dimensions. Instead of a line of best fit, it’s a plane of best fit:
Warning: Unknown or uninitialised column: `color`.bm_fl_bl_fit <- linear_reg() |>
fit(body_mass_g ~ flipper_length_mm + bill_length_mm, data = penguins)
tidy(bm_fl_bl_fit)# A tibble: 3 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) -5737. 308. -18.6 7.80e-54
2 flipper_length_mm 48.1 2.01 23.9 7.56e-75
3 bill_length_mm 6.05 5.18 1.17 2.44e- 1\[ \small\widehat{body~mass}=-5736+48.1\times flipper~length+6\times bill~length \]
The code was basically the same every time. In this second half of the course, the code itself is not the primary complication. The code is merely a labor-saving device. Our focus shifts to understanding the conceptual and technical underpinnings of the code and interpreting the output correctly. This is subtle, and it tends to trip people up.
Regarding the leveling of a categorical predictor:
Sort of. You get the same estimates of the slopes and intercepts, but the uncertainty quantification (standard errors) is different.
This matters, but we suppress the nuance in this class.
linear_reg() |>
fit(body_mass_g ~ flipper_length_mm * island, data = penguins) |>
tidy() |>
select(term, estimate, std.error)# A tibble: 6 × 3
term estimate std.error
<chr> <dbl> <dbl>
1 (Intercept) -5464. 431.
2 flipper_length_mm 48.5 2.05
3 islandDream 3551. 969.
4 islandTorgersen 3218. 1680.
5 flipper_length_mm:islandDream -19.4 4.94
6 flipper_length_mm:islandTorgersen -17.4 8.73So on Biscoe island (where Dream = Torgersen = 0), the line is
\[ \begin{aligned} \widehat{body~mass} = -5464 &+ 48.5 \times flipper~length \end{aligned} \]
If we fit a simple model with the Biscoe subset of the data:
penguins_biscoe <- penguins |>
filter(island == "Biscoe")
linear_reg() |>
fit(body_mass_g ~ flipper_length_mm, data = penguins_biscoe) |>
tidy()# A tibble: 2 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) -5464. 435. -12.6 8.48e-26
2 flipper_length_mm 48.5 2.07 23.4 2.20e-54Same estimates, but different standard errors.
ggplot(penguins, aes(x = flipper_length_mm, y = body_mass_g, color = island)) +
geom_point() +
geom_smooth(method = "lm")`geom_smooth()` using formula = 'y ~ x'Warning: Removed 2 rows containing non-finite outside the scale range
(`stat_smooth()`).Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_point()`).These are separate simple regressions within each group. The slopes and the intercepts are the same as the interaction model, but the uncertainty bands are different.
\(R^2\) measures “goodness-of-fit:”
So what is it, actually?
Every data point has one:
Some are big, some are small. Some are above the line (positive), and some are below the line (negative).
The residuals are what’s left over after the model attempts to “explain” the response:
\[ e_i = \text{observed} - \text{predicted} = y_i - \hat{y}_i. \]
Every data point has one:
# A tibble: 344 × 4
flipper_length_mm body_mass_g .pred .resid
<int> <int> <dbl> <dbl>
1 181 3750 3212. 538.
2 186 3800 3461. 339.
3 195 3250 3908. -658.
4 NA NA NA NA
5 193 3450 3808. -358.
6 190 3650 3659. -9.43
7 181 3625 3212. 413.
8 195 4675 3908. 767.
9 193 3475 3808. -333.
10 190 4250 3659. 591.
# ℹ 334 more rowsThe original distribution of the response. This is the mess we’re trying to clean up with a model:
# A tibble: 1 × 3
type variance std
<chr> <dbl> <dbl>
1 body_mass_g 643131. 802.The distribution of the residuals (the leftover mess) after the model tries to explain (clean up):
# A tibble: 2 × 3
type variance std
<chr> <dbl> <dbl>
1 .resid 154999. 394.
2 body_mass_g 643131. 802.The model is trying to “explain” the variation in the response:
That’s what \(R^2\) actually is: the proportion of the variance in the response explained by the model.
Horrifically awful fit. The model didn’t explain (clean up) anything:
# A tibble: 1 × 3
var_y var_resid r.squared
<dbl> <dbl> <dbl>
1 1.02 1.02 0.000450# A tibble: 1 × 3
var_y var_resid r.squared
<dbl> <dbl> <dbl>
1 0.805 0.574 0.287# A tibble: 1 × 3
var_y var_resid r.squared
<dbl> <dbl> <dbl>
1 0.701 0.255 0.636# A tibble: 1 × 3
var_y var_resid r.squared
<dbl> <dbl> <dbl>
1 0.762 0.0230 0.970Perfect fit. The model explained (cleaned up) everything:
# A tibble: 1 × 3
var_y var_resid r.squared
<dbl> <dbl> <dbl>
1 0.847 0.000408 1.000\[ \begin{aligned} R^2\, = \begin{matrix} \text{proportion of}\\ \text{variation explained} \end{matrix} &= 1- \begin{matrix} \text{proportion of}\\ \text{variation unexplained} \end{matrix}\\[15pt] &= 1-\frac{\text{unexplained variation}}{\text{total variation}} \\[15pt] &= 1-\frac{\text{spread of residuals}}{\text{spread of response}} \\[15pt] &= 1-\frac{\text{var}(e_i)}{\text{var}(y_i)} . \end{aligned} \]
\(R^2\) is a number between zero and one, and it represents the proportion of the variance in the response \(Y\) that is explained by the model fit;
This is the interpretation, regardless how many predictors you have;
In simple regression with one numerical predictor, \(R^2\) it is also the square of the correlation coefficient, but this doesn’t generalize to the multivariate setting.
Question: Given many competing models for the same response, which model is “best”? What does that even mean?
Answer: Assign a “quality score” to each model, and pick the model with the best score:
| Model | Score | Verdict |
|---|---|---|
| Simple | 0.1 | ❌ |
| Additive | 0.5 | ✅ |
| Interaction | 0.48 | ❌ |
Variable selection: all of our models were linear, and they differed only by which predictor variables are included. By ranking models by quality score and picking the “best” one, we can determine which predictors are the “most important” to include.
This is a big area of statistical research. There are loads of “quality scores” you could consider, and they prioritize different things: predictive accuracy, goodness-of-fit, simplicity, fairness, etc.
We won’t go into detail, but you will if you keep studying statistics.
tidymodels reports many of these for free\(R^2\) gives participation trophies. It always goes up when you add any new variable to the model, even if that variable is worthless.
The original model fit:
[1] 0.7589925Because vanilla \(R^2\) blithely rewards the addition of any variable, no matter how meaningless, you shouldn’t use it for model selection. Instead, use adjusted \(R^2\). The formula is in the textbook if you’re dying to know, but here’s the tea:
More complex models (i.e., models with more predictors) tend to fit the data at hand better, but may not generalize well to new data.
Model selection criteria, like adjusted \(R^2\), help balance model fit and complexity to avoid overfitting by penalizing models with more predictors.
Overfitting occurs when a model captures not only the underlying relationship between predictors and outcome but also the random noise in the data.
Overfitted models tend to perform well on the observed data but poorly on new, unseen data.