AE 23: Pokemon Inference
Today we’ll explore the question “Are taller Pokémon speedier or faster than shorter Pokémon?”
Packages
Data
The data for this application exercise was originally gathered using the Complete Pokémon Dataset from Kaggle.
First, let’s load the data:
pokemon <- read_csv("data/pokemon.csv")To keep things simple, we’ll work with a subset of the data, the generation 4 Pokémon.
Rows: 121
Columns: 17
$ name <chr> "Turtwig", "Grotle", "Torterra", "Chimchar", "Monferno…
$ generation <dbl> 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, …
$ type_1 <chr> "Grass", "Grass", "Grass", "Fire", "Fire", "Fire", "Wa…
$ type_2 <chr> NA, NA, "Ground", NA, "Fighting", "Fighting", NA, NA, …
$ height_m <dbl> 0.4, 1.1, 2.2, 0.5, 0.9, 1.2, 0.4, 0.8, 1.7, 0.3, 0.6,…
$ weight_kg <dbl> 10.2, 97.0, 310.0, 6.2, 22.0, 55.0, 5.2, 23.0, 84.5, 2…
$ total_points <dbl> 318, 405, 525, 309, 405, 534, 314, 405, 530, 245, 340,…
$ hp <dbl> 55, 75, 95, 44, 64, 76, 53, 64, 84, 40, 55, 85, 59, 79…
$ attack <dbl> 68, 89, 109, 58, 78, 104, 51, 66, 86, 55, 75, 120, 45,…
$ defense <dbl> 64, 85, 105, 44, 52, 71, 53, 68, 88, 30, 50, 70, 40, 6…
$ sp_attack <dbl> 45, 55, 75, 58, 78, 104, 61, 81, 111, 30, 40, 50, 35, …
$ sp_defense <dbl> 55, 65, 85, 44, 52, 71, 56, 76, 101, 30, 40, 60, 40, 6…
$ speed <dbl> 31, 36, 56, 61, 81, 108, 40, 50, 60, 60, 80, 100, 31, …
$ catch_rate <dbl> 45, 45, 45, 45, 45, 45, 45, 45, 45, 255, 120, 45, 255,…
$ base_friendship <dbl> 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70…
$ base_experience <dbl> 64, 142, 236, 62, 142, 240, 63, 142, 239, 49, 119, 218…
$ is_legendary <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, FALSE…We are interested in whether a Pokémon’s log height (measured in meters) can predict its speed (how fast it is).
Visualize
- Review! Create a new variable for the Pokémon’s log height.
- Your turn: Plot the data and the line of best fit.
ggplot(pokemon, aes(x = log_height, y = speed)) +
geom_point() +
geom_smooth(method = "lm")
Point Estimation
- Your turn: Fit the linear model to these data:
observed_fit <- pokemon |>
specify(speed ~ log_height) |>
fit()
observed_fit# A tibble: 2 × 2
term estimate
<chr> <dbl>
1 intercept 72.4
2 log_height 7.39This gives the exact same numbers that you get if you use linear_reg() |> fit(), but we need this new syntax because it plays nice with the tools we have for confidence intervals and hypothesis tests. I know, I hate it too, but it’s the way it is.
- Your turn: Typeset the equation for the model fit:
\[ \widehat{\text{speed}} = 72.41 + 7.39\times \log(\text{height}) \]
-
Your turn: Interpret the slope and the intercept estimates:
- If a Pokémon had 0 log(height) (equivalently 1 m tall), we predict that it would have a 72.41 speed stat on average;
- A $1 increase in log(height) is associated with a 7.39 increase in speed, on average.
Interval Estimation
-
Demo: Using seed
24601, generate500bootstrap samples, and store them in a new data frame calledbstrap_samples.
set.seed(24601)
bstrap_samples <- pokemon |>
specify(speed ~ log_height) |>
generate(reps = 500, type = "bootstrap")-
Demo: Fit a linear model to each of these bootstrap samples and store the estimates in a new data framed called
bstrap_fits.
bstrap_fits <- bstrap_samples |>
fit()-
Your turn: Use
linear_reg() |> fit(...)to fit a linear model to bootstrap sample number 347, and verify that you get the same estimates as the ones contained inbstrap_fits.
replicate_347 <- bstrap_samples |>
filter(
replicate == 347
)
linear_reg() |>
fit(speed ~ log_height, data = replicate_347) |>
tidy()# A tibble: 2 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 71.6 2.61 27.4 2.64e-53
2 log_height 6.46 3.09 2.09 3.84e- 2bstrap_fits |>
filter(replicate == 347)# A tibble: 2 × 3
# Groups: replicate [1]
replicate term estimate
<int> <chr> <dbl>
1 347 intercept 71.6
2 347 log_height 6.46The only point I’m making here is that this new bootstrap code is not performing a fundamentally new task. It’s performing an old task (fitting the linear model), but it’s repeating it A LOT. So the numbers you get here are not mysterious. They’re numbers you already know how to compute.
-
Demo: Compute 95% confidence intervals for the slope and the intercept using the
get_confidence_intervalcommand.
ci_95 <- get_confidence_interval(
bstrap_fits,
point_estimate = observed_fit,
level = 0.90,
type = "percentile"
)
ci_95# A tibble: 2 × 3
term lower_ci upper_ci
<chr> <dbl> <dbl>
1 intercept 68.2 76.3
2 log_height 2.22 12.4-
Your turn: Verify that you get the same numbers when you manually calculate the quantiles of the slope estimates using
summarizeandquantile. Pay attention to the grouping.
bstrap_fits |>
ungroup() |>
group_by(term) |>
summarize(
lower_ci = quantile(estimate, 0.05),
upper_ci = quantile(estimate, 0.95)
)# A tibble: 2 × 3
term lower_ci upper_ci
<chr> <dbl> <dbl>
1 intercept 68.2 76.3
2 log_height 2.22 12.4Same point as before. There’s no magic here. get_confidence_interval is just a convenient way of doing something that you already knew how to do.s
- BONUS: You can visualize the confidence interval:
visualize(bstrap_fits) +
shade_confidence_interval(ci_95)
Hypothesis Testing
Let’s consider the hypotheses:
\[ H_0:\beta_1=0\quad vs\quad H_A: \beta_1\neq 0. \] The null hypothesis corresponds to the claim that log_height and speed are uncorrelated.
- Simulate and plot the null distribution for the slope:
set.seed(24601)
null_dist <- pokemon |>
specify(speed ~ log_height) |>
hypothesize(null = "independence") |>
generate(reps = 500, type = "permute") |>
fit()
null_dist |>
filter(term == "log_height") |>
ggplot(aes(x = estimate)) +
geom_histogram()
- Add a vertical line to your plot indicating the point estimate of the slope from your original data analysis:
visualize(null_dist) +
shade_p_value(obs_stat = observed_fit, direction = "two-sided")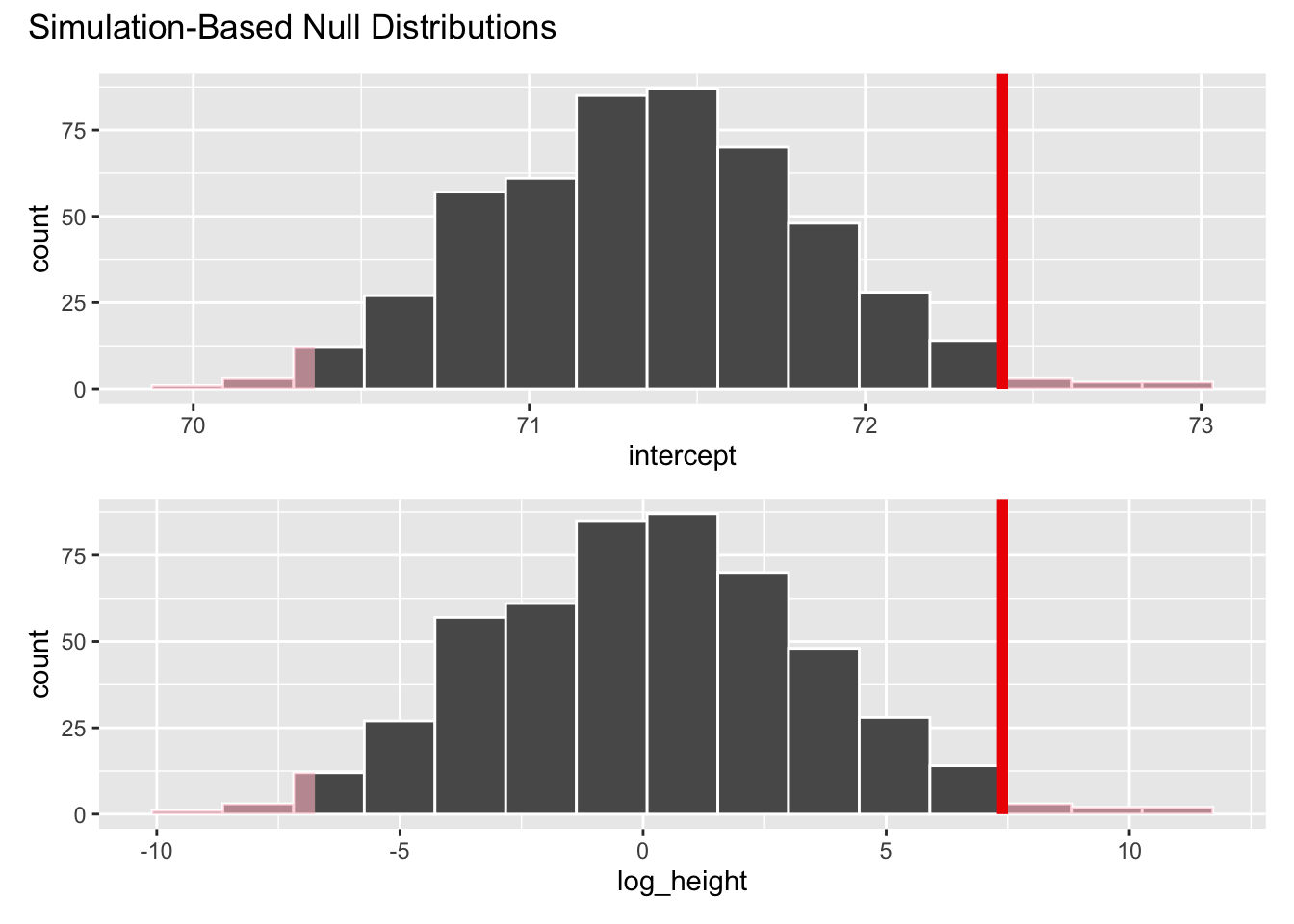
- Compute the \(p\)-value for this test and interpret it:
null_dist |>
get_p_value(obs_stat = observed_fit, direction = "two-sided")# A tibble: 2 × 2
term p_value
<chr> <dbl>
1 intercept 0.028
2 log_height 0.028Answer: If the null were true and \(\beta_1=0\) (meaning log_height and speed are uncorrelated), the probability of getting an estimate as or more extreme than the one we got (\(b_1=7.39\)) is 0.028. So we reject the null hypothesis. Evidence is sufficient to conclude that \(\beta_1\), whatever it is, is not equal to zero.
